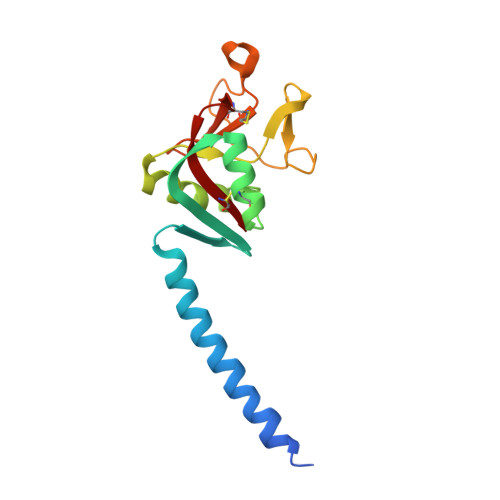
Contributions of Phenylalanine 335 to Ligand Recognition by Human Surfactant Protein D: ring interactions with Sp-D ligands
Crouch, E., McDonald, B., Smith, K., Cafarella, T., Seaton, B., Head, J.(2006) J Biological Chem 281: 18008-18014
- PubMed: 16636058 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M601749200
- Primary Citation Related Structures:
2GGU, 2GGX - PubMed Abstract:
Surfactant protein D (SP-D) is an innate immune effector that contributes to antimicrobial host defense and immune regulation. Interactions of SP-D with microorganisms and organic antigens involve binding of glycoconjugates to the C-type lectin carbohydrate recognition domain (CRD). A trimeric fusion protein encoding the human neck+CRD bound to the aromatic glycoside p-nitrophenyl-alpha-D-maltoside with nearly a log-fold higher affinity than maltose, the prototypical competitor. Maltotriose, which has the same linkage pattern as the maltoside, bound with intermediate affinity. Site-directed substitution of leucine for phenylalanine 335 (Phe-335) decreased affinities for the maltoside and maltotriose without significantly altering the affinity for maltose or glucose, and substitution of tyrosine or tryptophan for leucine restored preferential binding to maltotriose and the maltoside. A mutant with alanine at this position failed to bind to mannan or maltose-substituted solid supports. Crystallographic analysis of the human neck+CRD complexed with maltotriose or p-nitrophenyl-maltoside showed stacking of the terminal glucose or nitrophenyl ring with the aromatic ring of Phe-335. Our studies indicate that Phe-335, which is evolutionarily conserved in all known SP-Ds, plays important, if not critical, roles in SP-D function.
- Department of Pathology and Immunology, Washington University School of Medicine, 660 S. Euclid Avenue, St. Louis, MO 63110, USA. crouch@path.wustl.edu
Organizational Affiliation: