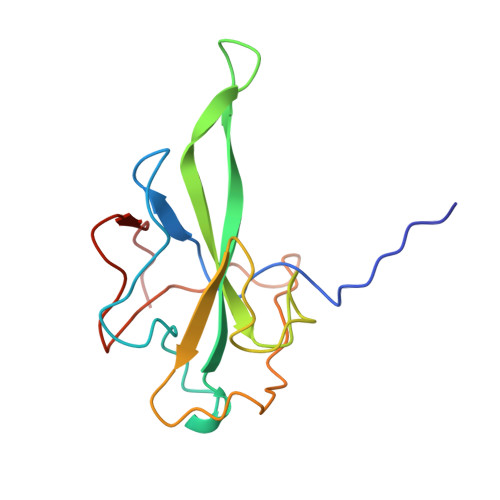
X-ray structures of the N- and C-terminal domains of a coronavirus nucleocapsid protein: implications for nucleocapsid formation.
Jayaram, H., Fan, H., Bowman, B.R., Ooi, A., Jayaram, J., Collisson, E.W., Lescar, J., Prasad, B.V.(2006) J Virol 80: 6612-6620
- PubMed: 16775348 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00157-06
- Primary Citation Related Structures:
2C86, 2CA1, 2GE7, 2GE8, 2GEC - PubMed Abstract:
Coronaviruses cause a variety of respiratory and enteric diseases in animals and humans including severe acute respiratory syndrome. In these enveloped viruses, the filamentous nucleocapsid is formed by the association of nucleocapsid (N) protein with single-stranded viral RNA. The N protein is a highly immunogenic phosphoprotein also implicated in viral genome replication and in modulating cell signaling pathways. We describe the structure of the two proteolytically resistant domains of the N protein from infectious bronchitis virus (IBV), a prototype coronavirus. These domains are located at its N- and C-terminal ends (NTD and CTD, respectively). The NTD of the IBV Gray strain at 1.3-A resolution exhibits a U-shaped structure, with two arms rich in basic residues, providing a module for specific interaction with RNA. The CTD forms a tightly intertwined dimer with an intermolecular four-stranded central beta-sheet platform flanked by alpha helices, indicating that the basic building block for coronavirus nucleocapsid formation is a dimeric assembly of N protein. The variety of quaternary arrangements of the NTD and CTD revealed by the analysis of the different crystal forms delineates possible interfaces that could be used for the formation of a flexible filamentous ribonucleocapsid. The striking similarity between the dimeric structure of CTD and the nucleocapsid-forming domain of a distantly related arterivirus indicates a conserved mechanism of nucleocapsid formation for these two viral families.
- Verna and Marrs McLean Department of Biochemistry and Molecular Biology, Baylor College of Medicine, One Baylor Plaza, Houston, TX 77030-3498, USA.
Organizational Affiliation: