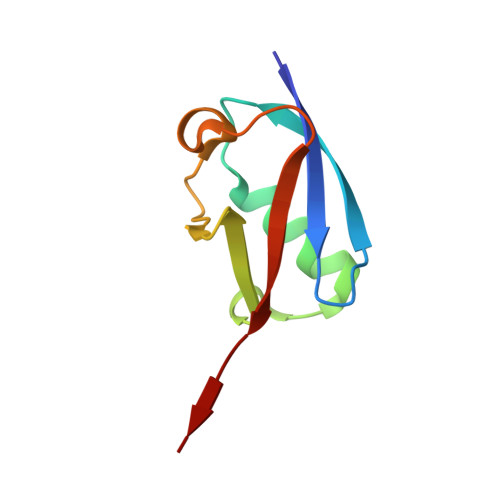
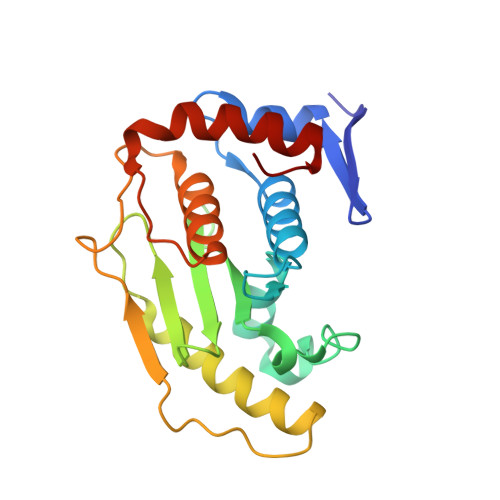
Crystal structure of the SENP1 mutant C603S-SUMO complex reveals the hydrolytic mechanism of SUMO-specific protease
Xu, Z., Chau, S.F., Lam, K.H., Chan, H.Y., Ng, T.B., Au, S.W.N.(2006) Biochem J 398: 345-352
- PubMed: 16712526 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20060526
- Primary Citation Related Structures:
2G4D - PubMed Abstract:
SUMO (small ubiquitin-related modifier)-specific proteases catalyse the maturation and de-conjugation processes of the sumoylation pathway and modulate various cellular responses including nuclear metabolism and cell cycle progression. The active-site cysteine residue is conserved among all known SUMO-specific proteases and is not substitutable by serine in the hydrolysis reactions demonstrated previously in yeast. We report here that the catalytic domain of human protease SENP1 (SUMO-specific protease 1) mutant SENP1C(C603S) carrying a mutation of cysteine to serine at the active site is inactive in maturation and de-conjugation reactions. To further understand the hydrolytic mechanism catalysed by SENP1, we have determined, at 2.8 A resolution (1 A = 0.1 nm), the X-ray structure of SENP1C(C603S)-SUMO-1 complex. A comparison of the structure of SENP2-SUMO-1 suggests strongly that SUMO-specific proteases require a self-conformational change prior to cleavage of peptide or isopeptide bond in the maturation and de-conjugation processes respectively. Moreover, analysis of the interface of SENP1 and SUMO-1 has led to the identification of four unique amino acids in SENP1 that facilitate the binding of SUMO-1. By means of an in vitro assay, we further demonstrate a novel function of SENP1 in hydrolysing the thioester linkage in E1-SUMO and E2-SUMO complexes. The results disclose a new mechanism of regulation of the sumoylation pathway by the SUMO-specific proteases.
- Department of Biochemistry and Molecular Biotechnology Program, Centre for Protein Science and Crystallography, Faculty of Science, The Chinese University of Hong Kong, Shatin, Hong Kong.
Organizational Affiliation: