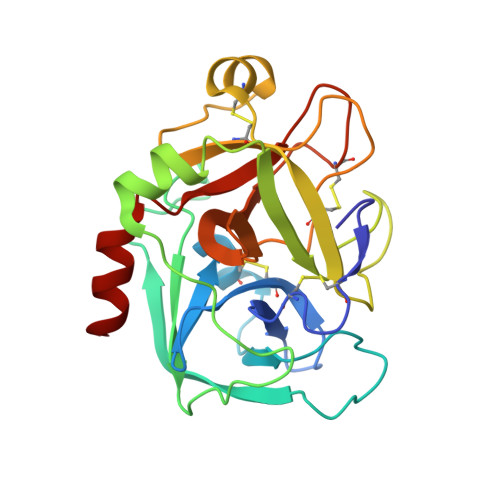
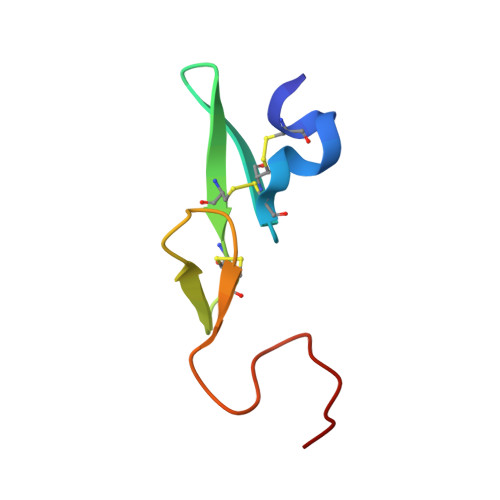
1-[3-Aminobenzisoxazol-5'-yl]-3-trifluoromethyl-6-[2'-(3-(R)-hydroxy-N-pyrrolidinyl)methyl-[1,1']-biphen-4-yl]-1,4,5,6-tetrahydropyrazolo-[3,4-c]-pyridin-7-one (BMS-740808) a highly potent, selective, efficacious, and orally bioavailable inhibitor of blood coagulation factor Xa.
Pinto, D.J., Orwat, M.J., Quan, M.L., Han, Q., Galemmo, R.A., Amparo, E., Wells, B., Ellis, C., He, M.Y., Alexander, R.S., Rossi, K.A., Smallwood, A., Wong, P.C., Luettgen, J.M., Rendina, A.R., Knabb, R.M., Mersinger, L., Kettner, C., Bai, S., He, K., Wexler, R.R., Lam, P.Y.(2006) Bioorg Med Chem Lett 16: 4141-4147
- PubMed: 16730984 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2006.02.069
- Primary Citation Related Structures:
2FZZ - PubMed Abstract:
Attempts to further optimize the pyrazole factor Xa inhibitors centered on masking the aryl aniline P4 moiety. Scaffold optimization resulted in the identification of a novel bicyclic pyrazolo-pyridinone scaffold which retained fXa potency. The novel bicyclic scaffold preserved all binding interactions observed with the monocyclic counterpart and importantly the carboxamido moiety was integrated within the scaffold making it less susceptible to hydrolysis. These efforts led to the identification of 1-[3-aminobenzisoxazol-5'-yl]-3-trifluoromethyl-6-[2'-(3-(R)-hydroxy-N-pyrrolidinyl)methyl-[1,1']-biphen-4-yl]-1,4,5,6-tetrahydropyrazolo-[3,4-c]-pyridin-7-one 6f (BMS-740808), a highly potent (fXa Ki=30 pM) with a rapid onset of inhibition (2.7x10(7) M-1 s-1) in vitro, selective (>1000-fold over other proteases), efficacious in the AVShunt thrombosis model, and orally bioavailable inhibitor of blood coagulation factor Xa.
- Discovery Chemistry Bristol-Myers Squibb Pharmaceutical Research Institute, Princeton, NJ 08543, USA. Donald.Pinto@bms.com
Organizational Affiliation: