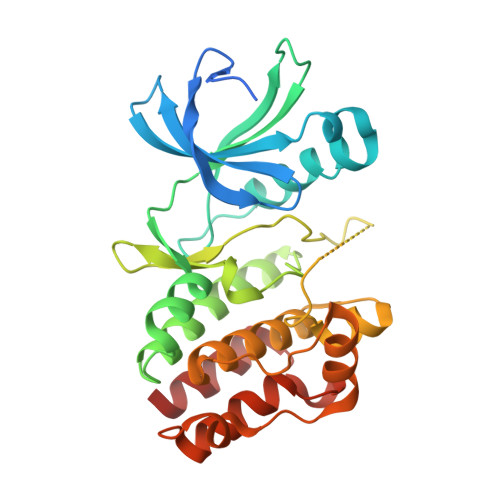
The structure of PknB in complex with mitoxantrone, an ATP-competitive inhibitor, suggests a mode of protein kinase regulation in mycobacteria
Wehenkel, A., Fernandez, P., Bellinzoni, M., Catherinot, V., Barilone, N., Labesse, G., Jackson, M., Alzari, P.M.(2006) FEBS Lett 580: 3018-3022
- PubMed: 16674948 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2006.04.046
- Primary Citation Related Structures:
2FUM - PubMed Abstract:
Mycobacterium tuberculosis PknB is an essential receptor-like protein kinase involved in cell growth control. Here, we demonstrate that mitoxantrone, an anthraquinone derivative used in cancer therapy, is a PknB inhibitor capable of preventing mycobacterial growth. The structure of the complex reveals that mitoxantrone partially occupies the adenine-binding pocket in PknB, providing a framework for the design of compounds with potential therapeutic applications. PknB crystallizes as a 'back-to-back' homodimer identical to those observed in other structures of PknB in complex with ATP analogs. This organization resembles that of the RNA-dependent protein kinase PKR, suggesting a mechanism for kinase activation in mycobacteria.
- Unité de Biochimie Structurale and CNRS URA2185, Institut Pasteur, 25 rue du Docteur Roux, 75724 Paris, France.
Organizational Affiliation: