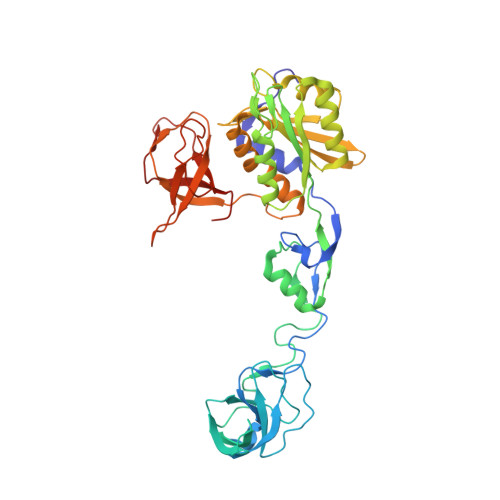
Deciphering the structural framework of glycine receptor anchoring by gephyrin.
Kim, E.Y., Schrader, N., Smolinsky, B., Bedet, C., Vannier, C., Schwarz, G., Schindelin, H.(2006) EMBO J 25: 1385-1395
- PubMed: 16511563 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601029
- Primary Citation Related Structures:
2FTS, 2FU3 - PubMed Abstract:
Glycine is the major inhibitory neurotransmitter in the spinal cord and brain stem. Gephyrin is required to achieve a high concentration of glycine receptors (GlyRs) in the postsynaptic membrane, which is crucial for efficient glycinergic signal transduction. The interaction between gephyrin and the GlyR involves the E-domain of gephyrin and a cytoplasmic loop located between transmembrane segments three and four of the GlyR beta subunit. Here, we present crystal structures of the gephyrin E-domain with and without the GlyR beta-loop at 2.4 and 2.7 A resolutions, respectively. The GlyR beta-loop is bound in a symmetric 'key and lock' fashion to each E-domain monomer in a pocket adjacent to the dimer interface. Structure-guided mutagenesis followed by in vitro binding and in vivo colocalization assays demonstrate that a hydrophobic interaction formed by Phe 330 of gephyrin and Phe 398 and Ile 400 of the GlyR beta-loop is crucial for binding.
- Department of Biochemistry, Center for Structural Biology, State University of New York, Stony Brook, NY 11794-5115, USA.
Organizational Affiliation: