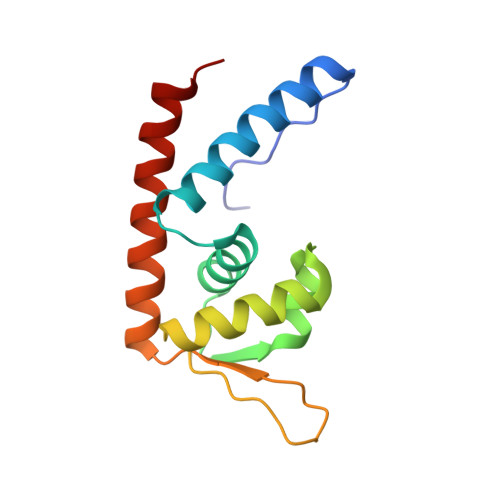
Structural and function analyses of the global regulatory protein SarA from Staphylococcus aureus.
Liu, Y., Manna, A.C., Pan, C.H., Kriksunov, I.A., Thiel, D.J., Cheung, A.L., Zhang, G.(2006) Proc Natl Acad Sci U S A 103: 2392-2397
- PubMed: 16455801
- DOI: https://doi.org/10.1073/pnas.0510439103
- Primary Citation Related Structures:
2FNP, 2FRH - PubMed Abstract:
The sarA locus in Staphylococcus aureus controls the expression of many virulence genes. The sarA regulatory molecule, SarA, is a 14.7-kDa protein (124 residues) that binds to the promoter region of target genes. Here we report the 2.6 A-resolution x-ray crystal structure of the dimeric winged helix SarA protein, which differs from the published SarA structure dramatically. In the crystal packing, multiple dimers of SarA form a scaffold, possibly via divalent cations. Mutations of individual residues within the DNA-binding helix-turn-helix and the winged region as well as within the metal-binding pocket implicate basic residues R84 and R90 within the winged region to be critical in DNA binding, whereas acidic residues D88 and E89 (wing), D8 and E11 (metal-binding pocket), and cysteine 9 are essential for SarA function. These data suggest that the winged region of the winged helix protein participates in DNA binding and activation, whereas the putative divalent cation binding pocket is only involved in gene function.
- Integrated Department of Immunology, National Jewish Medical and Research Center, Biomolecular Structure Program and Department of Pharmacology, School of Medicine, University of Colorado Health Science Center, Denver, CO 80206, USA.
Organizational Affiliation: