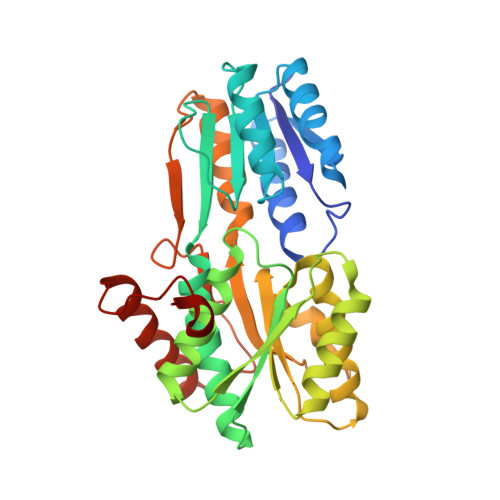
The PnrA (Tp0319; TmpC) lipoprotein represents a new family of bacterial purine nucleoside receptor encoded within an ATP-binding cassette (ABC)-like operon in Treponema pallidum
Deka, R.K., Brautigam, C.A., Yang, X.F., Blevins, J.S., Machius, M., Tomchick, D.R., Norgard, M.V.(2006) J Biological Chem 281: 8072-8081
- PubMed: 16418175
- DOI: https://doi.org/10.1074/jbc.M511405200
- Primary Citation of Related Structures:
2FQW, 2FQX, 2FQY - PubMed Abstract:
Treponema pallidum, the bacterial agent of syphilis, cannot be cultivated in vitro. This constraint has severely impeded the study of the membrane biology of this complex human pathogen. A structure-to-function approach thus was adopted as a means of discerning the likely function of Tp0319, a 35-kDa cytoplasmic membrane-associated lipoprotein of T. pallidum formerly designated as TmpC. A 1.7-A crystal structure showed that recombinant Tp0319 (rTp0319) consists of two alpha/beta domains, linked by three crossovers, with a deep cleft between them akin to ATP-binding cassette (ABC) receptors. In the cleft, a molecule of inosine was bound. Isothermal titration calorimetry demonstrated that rTp0319 specifically binds purine nucleosides (dissociation constant (Kd) approximately 10(-7) M). This predilection for purine nucleosides by rTp0319 is consistent with its likely role as a receptor component of a cytoplasmic membrane-associated transporter system. Reverse transcription-PCR analysis of RNA isolated from rabbit tissue-extracted T. pallidum additionally showed that tp0319 is transcriptionally linked to four other downstream open reading frames, thereby supporting the existence of an ABC-like operon (tp0319-0323). We herein thus re-name tp0319 as purine nucleoside receptor A (pnrA), with its operonic partners tp0320-0323 designated as pnrB-E, respectively. Our study not only infers that PnrA transports purine nucleosides essential for the survival of T. pallidum within its obligate human host, but to our knowledge, this is the first description of an ABC-type nucleoside transport system in any bacterium. PnrA has been grouped with a functionally uncharacterized protein family (HBG016869), thereby implying that other members of the family may have similar nucleoside-binding function(s).
- Department of Microbiology, University of Texas Southwestern Medical Center, Dallas, Texas 75390, USA.
Organizational Affiliation: