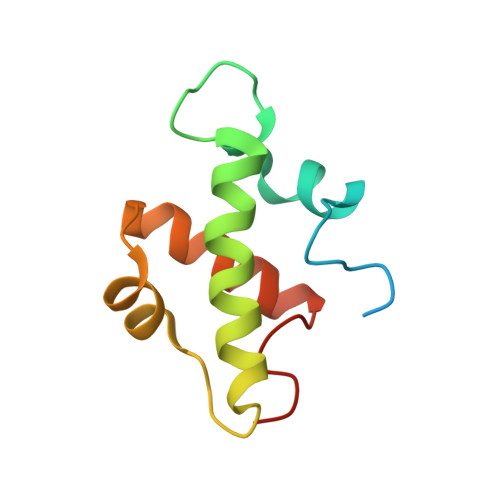
Structure and function of the SWIRM domain, a conserved protein module found in chromatin regulatory complexes
Da, G., Lenkart, J., Zhao, K., Shiekhattar, R., Cairns, B.R., Marmorstein, R.(2006) Proc Natl Acad Sci U S A 103: 2057-2062
- PubMed: 16461455 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0510949103
- Primary Citation Related Structures:
2FQ3 - PubMed Abstract:
The SWIRM domain is a module found in the Swi3 and Rsc8 subunits of SWI/SNF-family chromatin remodeling complexes, and the Ada2 and BHC110/LSD1 subunits of chromatin modification complexes. Here we report the high-resolution crystal structure of the SWIRM domain from Swi3 and characterize the in vitro and in vivo function of the SWIRM domains from Saccharomyces cerevisiae Swi3 and Rsc8. The Swi3 SWIRM forms a four-helix bundle containing a pseudo 2-fold axis and a helix-turn-helix motif commonly found in DNA-binding proteins. We show that the Swi3 SWIRM binds free DNA and mononucleosomes with high and comparable affinity and that a subset of Swi3 substitution mutants that display growth defects in vivo also show impaired DNA-binding activity in vitro, consistent with a nucleosome targeting function of this domain. Genetic and biochemical studies also reveal that the Rsc8 and Swi3 SWIRM domains are essential for the proper assembly and in vivo functions of their respective complexes. Together, these studies identify the SWIRM domain as an essential multifunctional module for the regulation of gene expression.
- The Wistar Institute and Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104, USA.
Organizational Affiliation: