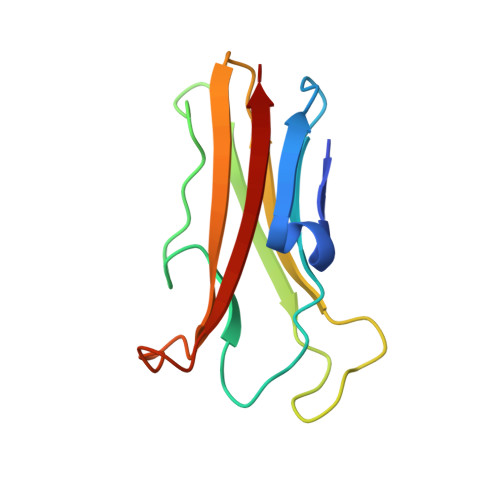
Solution Structure and Backbone Dynamics of the Trypanosoma cruzi Cysteine Protease Inhibitor Chagasin
Salmon, D., do Aido-Machado, R., Diehl, A., Leidert, M., Schmetzer, O., de Lima, A.A.P., Scharfstein, J., Oschkinat, H., Pires, J.R.(2006) J Mol Biology 357: 1511-1521
- PubMed: 16490204
- DOI: https://doi.org/10.1016/j.jmb.2006.01.064
- Primary Citation of Related Structures:
2FO8 - PubMed Abstract:
A Trypanosoma cruzi cysteine protease inhibitor, termed chagasin, is the first characterized member of a new family of tight-binding cysteine protease inhibitors identified in several lower eukaryotes and prokaryotes but not present in mammals. In the protozoan parasite T.cruzi, chagasin plays a role in parasite differentiation and in mammalian host cell invasion, due to its ability to modulate the endogenous activity of cruzipain, a lysosomal-like cysteine protease. In the present work, we determined the solution structure of chagasin and studied its backbone dynamics by NMR techniques. Structured as a single immunoglobulin-like domain in solution, chagasin exerts its inhibitory activity on cruzipain through conserved residues placed in three loops in the same side of the structure. One of these three loops, L4, predicted to be of variable length among chagasin homologues, is flexible in solution as determined by measurements of (15)N relaxation. The biological implications of structural homology between chagasin and other members of the immunoglobulin super-family are discussed.
- Instituto de Bioquímica Médica, CCS, Universidade Federal do Rio de Janeiro, Av. Brigadeiro Trompowiski s/n, Rio de Janeiro, RJ 21941-590, Brazil.
Organizational Affiliation: