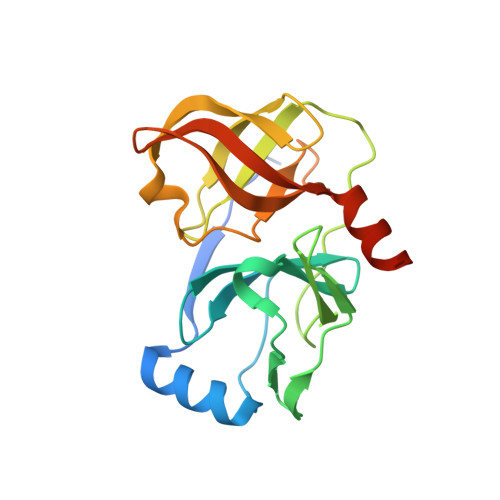
Mutations conferring resistance to SCH6, a novel hepatitis C virus NS3/4A protease inhibitor. Reduced RNA replication fitness and partial rescue by second-site mutations
Yi, M., Tong, X., Skelton, A., Chase, R., Chen, T., Prongay, A., Bogen, S.L., Saksena, A.K., Njoroge, F.G., Veselenak, R.L., Pyles, R.B., Bourne, N., Malcolm, B.A., Lemon, S.M.(2006) J Biological Chem 281: 8205-8215
- PubMed: 16352601
- DOI: https://doi.org/10.1074/jbc.M510246200
- Primary Citation Related Structures:
2FM2 - PubMed Abstract:
Drug resistance is a major issue in the development and use of specific antiviral therapies. Here we report the isolation and characterization of hepatitis C virus RNA replicons resistant to a novel ketoamide inhibitor of the NS3/4A protease, SCH6 (originally SCH446211). Resistant replicon RNAs were generated by G418 selection in the presence of SCH6 in a dose-dependent fashion, with the emergence of resistance reduced at higher SCH6 concentrations. Sequencing demonstrated remarkable consistency in the mutations conferring SCH6 resistance in genotype 1b replicons derived from two different strains of hepatitis C virus, A156T/A156V and R109K. R109K, a novel mutation not reported previously to cause resistance to NS3/4A inhibitors, conferred moderate resistance only to SCH6. Structural analysis indicated that this reflects unique interactions of SCH6 with P'-side residues in the protease active site. In contrast, A156T conferred high level resistance to SCH6 and a related ketoamide, SCH503034, as well as BILN 2061 and VX-950. Unlike R109K, which had minimal impact on NS3/4A enzymatic function, A156T significantly reduced NS3/4A catalytic efficiency, polyprotein processing, and replicon fitness. However, three separate second-site mutations, P89L, Q86R, and G162R, were capable of partially reversing A156T-associated defects in polyprotein processing and/or replicon fitness, without significantly reducing resistance to the protease inhibitor.
- Center for Hepatitis Research, Institute for Human Infections & Immunity, University of Texas Medical Branch, Galveston 77555-1019, USA.
Organizational Affiliation: