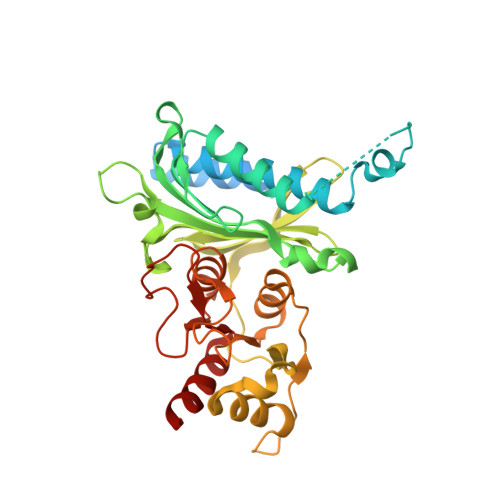
Benzoxazole benzenesulfonamides are novel allosteric inhibitors of fructose-1,6-bisphosphatase with a distinct binding mode.
von Geldern, T.W., Lai, C., Gum, R.J., Daly, M., Sun, C., Fry, E.H., Abad-Zapatero, C.(2006) Bioorg Med Chem Lett 16: 1811-1815
- PubMed: 16442285 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2006.01.015
- Primary Citation Related Structures:
2FHY - PubMed Abstract:
We have identified benzoxazole benzenesulfonamide 1 as a novel allosteric inhibitor of fructose-1,6-bisphosphatase (FBPase-1). X-ray crystallographic and biological studies of 1 indicate a distinct binding mode that recapitulates features of several previously reported FBPase-1 inhibitor classes.
- Metabolic Disease Research, GPRD, Abbott Laboratories, Abbott Park, IL 60064, USA. Thomas.vongeldern@abbott.com
Organizational Affiliation: