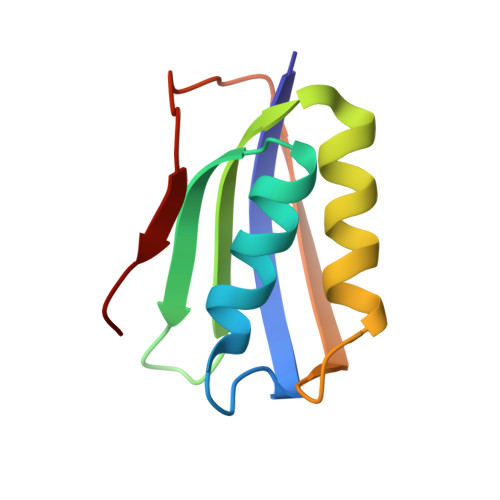
Solution structure and conformational heterogeneity of acylphosphatase from Bacillus subtilis
Hu, J.C., Li, D., Su, X.-D., Jin, C.W., Xia, B.(2010) FEBS Lett 584: 2852-2856
- PubMed: 20447399
- DOI: https://doi.org/10.1016/j.febslet.2010.04.069
- Primary Citation of Related Structures:
2FHM - PubMed Abstract:
Acylphosphatase is a small enzyme that catalyzes the hydrolysis of acyl phosphates. Here, we present the solution structure of acylphosphatase from Bacillus subtilis (BsAcP), the first from a Gram-positive bacterium. We found that its active site is disordered, whereas it converted to an ordered state upon ligand binding. The structure of BsAcP is sensitive to pH and it has multiple conformations in equilibrium at acidic pH (pH<5.8). Only one main conformation could bind ligand, and the relative population of these states is modulated by ligand concentration. This study provides direct evidence for the role of ligand in conformational selection.
- Beijing Nuclear Magnetic Resonance Center, Peking University, Beijing, China.
Organizational Affiliation: