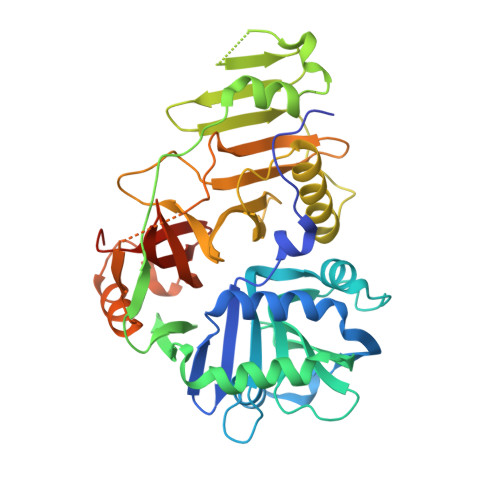
The crystal structure of YycH involved in the regulation of the essential YycFG two-component system in Bacillus subtilis reveals a novel tertiary structure.
Szurmant, H., Zhao, H., Mohan, M.A., Hoch, J.A., Varughese, K.I.(2006) Protein Sci 15: 929-934
- PubMed: 16600972 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.052064406
- Primary Citation Related Structures:
2FGT - PubMed Abstract:
The Bacillus subtilis YycFG two-component signal transduction system is essential for cell viability, and the YycH protein is part of the regulatory circuit that controls its activity. The crystal structure of YycH was solved by two-wavelength selenium anomalous dispersion data, and was refined using 2.3 A data to an R-factor of 25.2%. The molecule is made up of three domains, and has a novel three-dimensional structure. The N-terminal domain features a calcium binding site and the central domain contains two conserved loop regions.
- Division of Cellular Biology, Department of Molecular and Experimental Medicine, The Scripps Research Institute, La Jolla, California 92037, USA.
Organizational Affiliation: