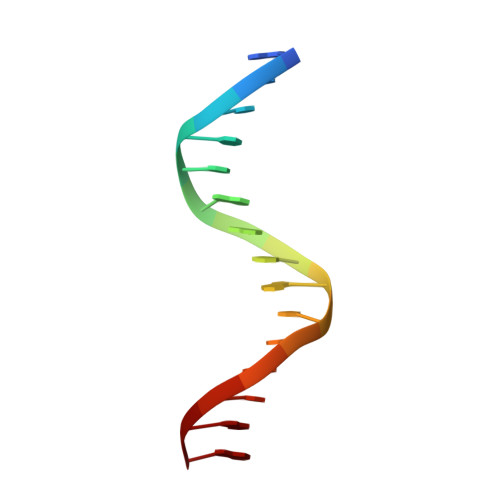
Sequence-specific deoxyribonucleic Acid (DNA) recognition by steroidogenic factor 1: a helix at the carboxy terminus of the DNA binding domain is necessary for complex stability.
Little, T.H., Zhang, Y., Matulis, C.K., Weck, J., Zhang, Z., Ramachandran, A., Mayo, K.E., Radhakrishnan, I.(2006) Mol Endocrinol 20: 831-843
- PubMed: 16339274
- DOI: https://doi.org/10.1210/me.2005-0384
- Primary Citation of Related Structures:
2FF0 - PubMed Abstract:
Steroidogenic factor 1 (SF1) is a member of the NR5A subfamily of nuclear hormone receptors and is considered a master regulator of reproduction because it regulates a number of genes encoding reproductive hormones and enzymes involved in steroid hormone biosynthesis. Like other NR5A members, SF1 harbors a highly conserved approximately 30-residue segment called the FTZ-F1 box C-terminal to the core DNA binding domain (DBD) common to all nuclear receptors and binds to 9-bp DNA sequences as a monomer. Here we describe the solution structure of the SF1 DBD in complex with an atypical sequence in the proximal promoter region of the inhibin-alpha gene that encodes a subunit of a reproductive hormone. SF1 forms a specific complex with the DNA through a bipartite motif binding to the major and minor grooves through the core DBD and the N-terminal segment of the FTZ-F1 box, respectively, in a manner previously described for two other monomeric receptors, nerve growth factor-induced-B and estrogen-related receptor 2. However, unlike these receptors, SF1 harbors a helix in the C-terminal segment of the FTZ-F1 box that interacts with both the core DBD and DNA and serves as an important determinant of stability of the complex. We propose that the FTZ-F1 helix along with the core DBD serves as a platform for interactions with coactivators and other DNA-bound factors in the vicinity.
- Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, Illinois 60208-3500, USA.
Organizational Affiliation: