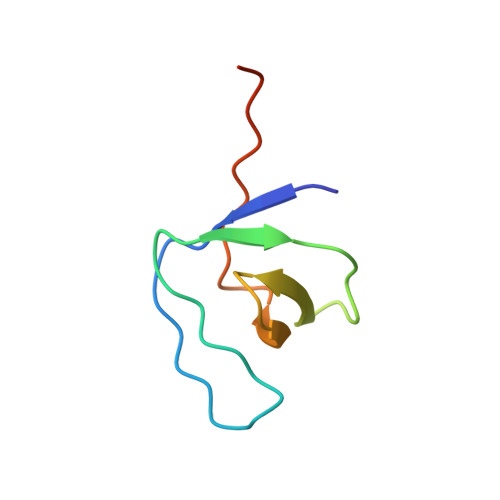
Solution structure of the second SH3 domain of human CMS and a newly identified binding site at the C-terminus of c-Cbl
Yao, B., Zhang, J., Dai, H., Sun, J., Jiao, Y., Tang, Y., Wu, J., Shi, Y.(2007) Biochim Biophys Acta 1774: 35-43
- PubMed: 17188587 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2006.09.018
- Primary Citation Related Structures:
2FEI - PubMed Abstract:
CMS, cas ligand with multiple Src homology 3 (SH3) domains, belongs to a family of ubiquitously expressed adaptor proteins. Among the CMS binding proteins, c-Cbl has been mostly extensively studied. It was reported that the motif PKPFPR (residues 824-829) of c-Cbl can bind to the N-terminus SH3 domains of CMS. Here we report the solution structure of the second SH3 domain of CMS (CMS_SH3_B), furthermore, we have identified that a peptide from residues 701 to 714 of c-Cbl (Cbl-p), i.e. MTPSSRPLRPLDTS, can specially bind to CMS_SH3_B using NMR chemical shift perturbation, suggesting that the peptide is a new potential CMS binding site. Among the peptide, TPSSRPLR is the core binding motif and Arg709 plays a key role in the interaction. Cbl-p binding interface on CMS_SH3_B along a hydrophobic channel is composed of RT loop, n-Src loop and beta4 strand and divided into three pockets. This work indicates the solution structure of CMS_SH3_B bears the canonical beta-beta-beta-beta-alpha-beta fold and a new binding site in c-Cbl involved in its interaction with CMS, which probably contributes to the clustering of CMS. All the information provided here should be beneficial for the future functional study of CMS.
- School of Life Science, Hefei National Laboratory for Physical Sciences at Microscale, University of Science and Technology of China, Hefei, Anhui 230027, Peoples Republic of China.
Organizational Affiliation: