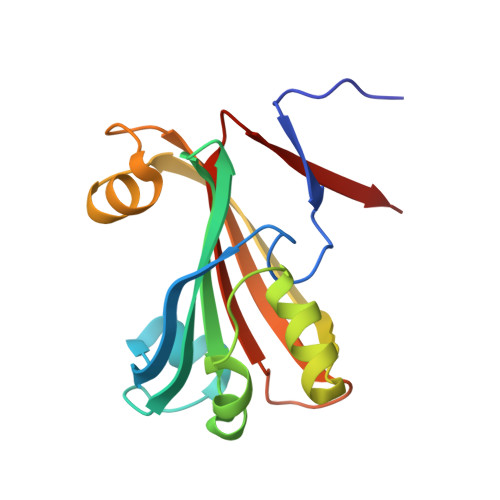
Structural characterization of GntR/HutC family signaling domain.
Gorelik, M., Lunin, V.V., Skarina, T., Savchenko, A.(2006) Protein Sci 15: 1-6
- PubMed: 16672238 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062146906
- Primary Citation Related Structures:
2FA1 - PubMed Abstract:
The crystal structure of Escherichia coli PhnF C-terminal domain (C-PhnF) was solved at 1.7 A resolution by the single wavelength anomalous dispersion (SAD) method. The PhnF protein belongs to the HutC subfamily of the large GntR transcriptional regulator family. Members of this family share similar N-terminal DNA-binding domains, but are divided into four subfamilies according to their heterogenic C-terminal domains, which are involved in effector binding and oligomerization. The C-PhnF structure provides for the first time the scaffold of this domain for the HutC subfamily, which covers about 31% of GntR-like regulators. The structure represents a mixture of alpha-helices and beta-strands, with a six-stranded antiparallel beta-sheet at the core. C-PhnF monomers form a dimer by establishing interdomain eight-strand beta-sheets that include core antiparallel and N-terminal two-strand parallel beta-sheets from each monomer. C-PhnF shares strong structural similarity with the chorismate lyase fold, which features a buried active site locked behind two helix-turn-helix loops. The structural comparison of the C-PhnF and UbiC proteins allows us to propose that a similar site in the PhnF structure is adapted for effector binding.
- Ontario Center for Structural Proteomics, University Health Network, University of Toronto, Toronto, Ontario M5G 1L7, Canada.
Organizational Affiliation: