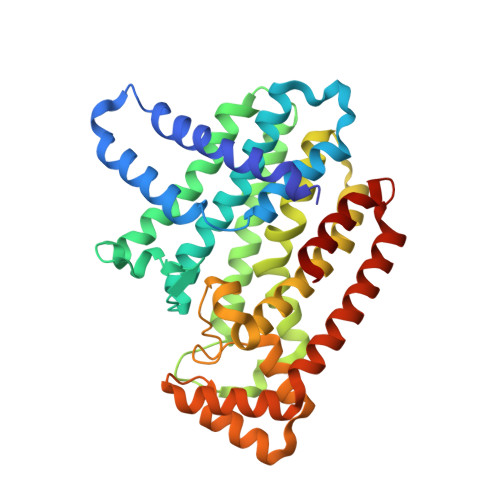
Structural basis for the exceptional in vivo efficacy of bisphosphonate drugs.
Rondeau, J.M., Bitsch, F., Bourgier, E., Geiser, M., Hemmig, R., Kroemer, M., Lehmann, S., Ramage, P., Rieffel, S., Strauss, A., Green, J.R., Jahnke, W.(2006) ChemMedChem 1: 267-273
- PubMed: 16892359
- DOI: https://doi.org/10.1002/cmdc.200500059
- Primary Citation Related Structures:
2F7M, 2F89, 2F8C, 2F8Z, 2F92, 2F94, 2F9K - PubMed Abstract:
To understand the structural basis for bisphosphonate therapy of bone diseases, we solved the crystal structures of human farnesyl pyrophosphate synthase (FPPS) in its unliganded state, in complex with the nitrogen-containing bisphosphonate (N-BP) drugs zoledronate, pamidronate, alendronate, and ibandronate, and in the ternary complex with zoledronate and the substrate isopentenyl pyrophosphate (IPP). By revealing three structural snapshots of the enzyme catalytic cycle, each associated with a distinct conformational state, and details about the interactions with N-BPs, these structures provide a novel understanding of the mechanism of FPPS catalysis and inhibition. In particular, the accumulating substrate, IPP, was found to bind to and stabilize the FPPS-N-BP complexes rather than to compete with and displace the N-BP inhibitor. Stabilization of the FPPS-N-BP complex through IPP binding is supported by differential scanning calorimetry analyses of a set of representative N-BPs. Among other factors such as high binding affinity for bone mineral, this particular mode of FPPS inhibition contributes to the exceptional in vivo efficacy of N-BP drugs. Moreover, our data form the basis for structure-guided design of optimized N-BPs with improved pharmacological properties.
- Novartis Institutes for BioMedical Research, Discovery Technologies, 4002 Basel, Switzerland. jeanmichel.rondeau@novartis.com
Organizational Affiliation: