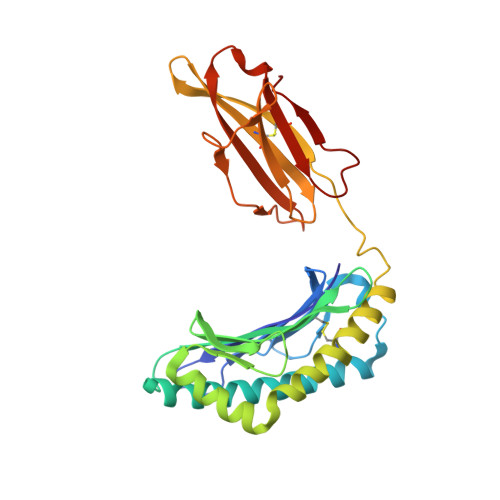
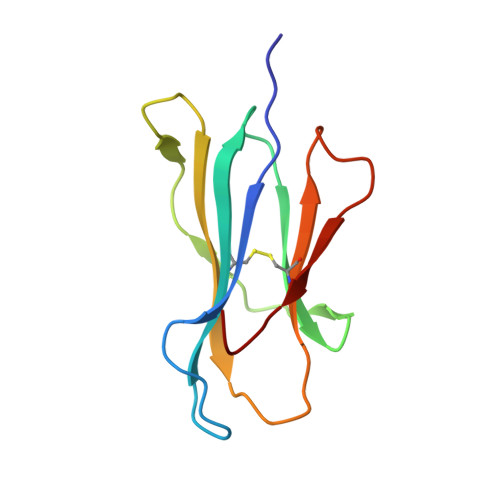
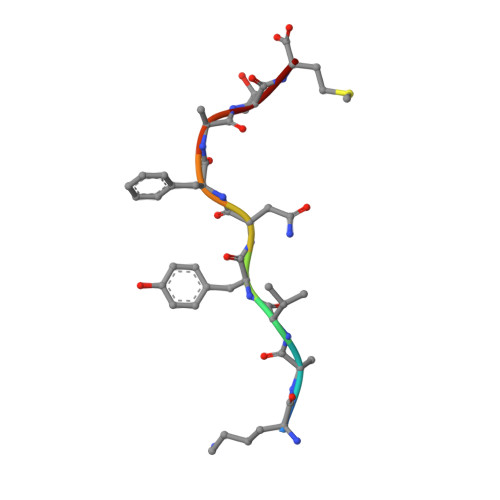
Structural Basis of the Differential Stability and Receptor Specificity of H-2D(b) in Complex with Murine versus Human beta(2)-Microglobulin.
Achour, A., Harris, R.A., Ljunggren, H.G., Schneider, G., Sandalova, T.(2006) J Mol Biology 356: 382-396
- PubMed: 16375919 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.11.068
- Primary Citation Related Structures:
2F74 - PubMed Abstract:
beta(2)-Microglobulin (beta(2)m) is non-covalently linked to the major histocompatibility complex (MHC) class I heavy chain and interacts with CD8 and Ly49 receptors. Murine MHC class I heavy chains can bind human beta(2)m (hbeta(2)m) and peptide, and such hybrid molecules are often used in structural and functional studies. The replacement of mouse beta(2)m (mbeta(2)m) with hbeta(2)m has several functional consequences for MHC class I complex stability and specificity, but the structural basis for this is presently unknown. To investigate the impact of species-specific beta(2)m subunits on MHC class I conformation, we provide a crystallographic comparison of H-2D(b) in complex with LCMV-derived gp33 peptide and either hbeta(2)m or mbeta(2)m. The conformation of the gp33 peptide is not affected by the beta(2)m species. Comparison of the interface between beta(2)m and the alpha(1)alpha(2) domains of the heavy chain in these two crystal structures reveals a marked increase in both polarity and number of hydrogen bonds between hbeta(2)m and the alpha(1)alpha(2) domains of H-2D(b). We propose that the positioning of two hydrogen bond rich regions at the hbeta(2)m/alpha(1)alpha(2) interface plays a central role in the increased overall stability and peptide exchange capacity in the H-2D(b)/hbeta(2)m complex. These two regions act as bridges, holding and stabilizing the underside of the alpha(1) and alpha(2) helices, enabling a prolonged peptide-receptive conformation of the peptide binding cleft. Furthermore, analysis of H-2D(b) in complex with either mbeta(2)m or hbeta(2)m provides a structural explanation for the differential binding of H-2D(b)/hbeta(2)m to both Ly49A and Ly49C. Our comparative structural study emphasizes the importance of beta(2)m residues at positions 3, 6 and 29 for binding to Ly49A and suggests that sterical hindrance by residue K6 on hbeta(2)m impairs the recognition of Ly49C by H-2D(b)/gp33/hbeta(2)m. Finally, comparison of the two H-2D(b) crystal structures implies that the beta(2)m species may affect the strength of TCR recognition by affecting CD8 binding.
- Center for Infectious Medicine, F59, Department of Medicine, Karolinska Institutet, Karolinska University Hospital in Huddinge, Stockholm, Sweden. adnane.achour@medjs.ki.se
Organizational Affiliation: