Biochemical and structural characterization of the secreted chorismate mutase (Rv1885c) from Mycobacterium tuberculosis H37Rv: an *AroQ enzyme not regulated by the aromatic amino acids.
Kim, S.K., Reddy, S.K., Nelson, B.C., Vasquez, G.B., Davis, A., Howard, A.J., Patterson, S., Gilliland, G.L., Ladner, J.E., Reddy, P.T.(2006) J Bacteriol 188: 8638-8648
- PubMed: 17146044 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00441-06
- Primary Citation Related Structures:
2F6L - PubMed Abstract:
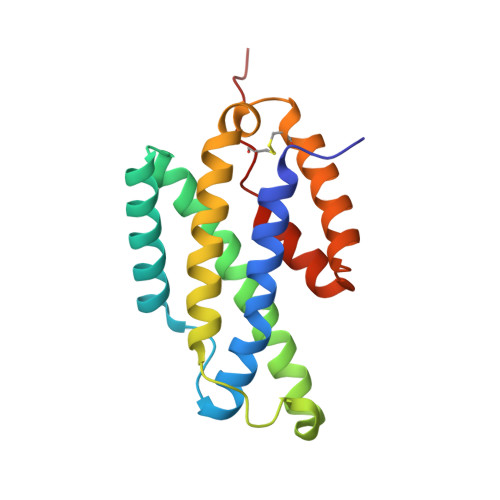
The gene Rv1885c from the genome of Mycobacterium tuberculosis H37Rv encodes a monofunctional and secreted chorismate mutase (*MtCM) with a 33-amino-acid cleavable signal sequence; hence, it belongs to the *AroQ class of chorismate mutases. Consistent with the heterologously expressed *MtCM having periplasmic destination in Escherichia coli and the absence of a discrete periplasmic compartment in M. tuberculosis, we show here that *MtCM secretes into the culture filtrate of M. tuberculosis. *MtCM functions as a homodimer and exhibits a dimeric state of the protein at a concentration as low as 5 nM. *MtCM exhibits simple Michaelis-Menten kinetics with a Km of 0.5 +/- 0.05 mM and a k(cat) of 60 s(-1) per active site (at 37 degrees C and pH 7.5). The crystal structure of *MtCM has been determined at 1.7 A resolution (Protein Data Bank identifier 2F6L). The protein has an all alpha-helical structure, and the active site is formed within a single chain without any contribution from the second chain in the dimer. Analysis of the structure shows a novel fold topology for the protein with a topologically rearranged helix containing Arg134. We provide evidence by site-directed mutagenesis that the residues Arg49, Lys60, Arg72, Thr105, Glu109, and Arg134 constitute the catalytic site; the numbering of the residues includes the signal sequence. Our investigation on the effect of phenylalanine, tyrosine, and tryptophan on *MtCM shows that *MtCM is not regulated by the aromatic amino acids. Consistent with this observation, the X-ray structure of *MtCM does not have an allosteric regulatory site.
- Biochemical Science Division, National Institute of Standards and Technology, Mail stop 831.2, Bldg. 227, Room B244, Gaithersburg, MD 20899, USA.
Organizational Affiliation: