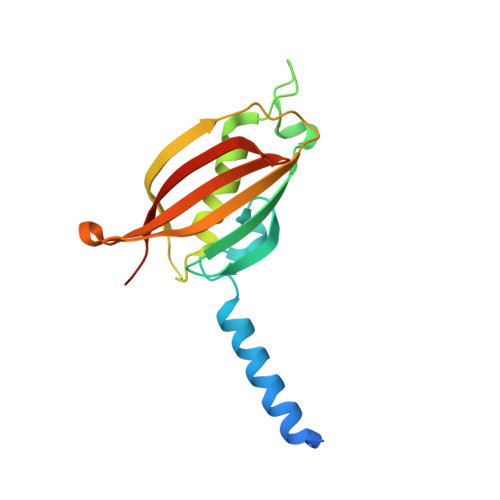
Structural basis of lipid biosynthesis regulation in Gram-positive bacteria.
Schujman, G.E., Guerin, M., Buschiazzo, A., Schaeffer, F., Llarrull, L.I., Reh, G., Vila, A.J., Alzari, P.M., de Mendoza, D.(2006) EMBO J 25: 4074-4083
- PubMed: 16932747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601284
- Primary Citation Related Structures:
2F3X, 2F41 - PubMed Abstract:
Malonyl-CoA is an essential intermediate in fatty acid synthesis in all living cells. Here we demonstrate a new role for this molecule as a global regulator of lipid homeostasis in Gram-positive bacteria. Using in vitro transcription and binding studies, we demonstrate that malonyl-CoA is a direct and specific inducer of Bacillus subtilis FapR, a conserved transcriptional repressor that regulates the expression of several genes involved in bacterial fatty acid and phospholipid synthesis. The crystal structure of the effector-binding domain of FapR reveals a homodimeric protein with a thioesterase-like 'hot-dog' fold. Binding of malonyl-CoA promotes a disorder-to-order transition, which transforms an open ligand-binding groove into a long tunnel occupied by the effector molecule in the complex. This ligand-induced modification propagates to the helix-turn-helix motifs, impairing their productive association for DNA binding. Structure-based mutations that disrupt the FapR-malonyl-CoA interaction prevent DNA-binding regulation and result in a lethal phenotype in B. subtilis, suggesting this homeostatic signaling pathway as a promising target for novel chemotherapeutic agents against Gram-positive pathogens.
- Instituto de Biología Molecular y Celular de Rosario (IBR), Universidad Nacional de Rosario, Rosario, Argentina.
Organizational Affiliation: