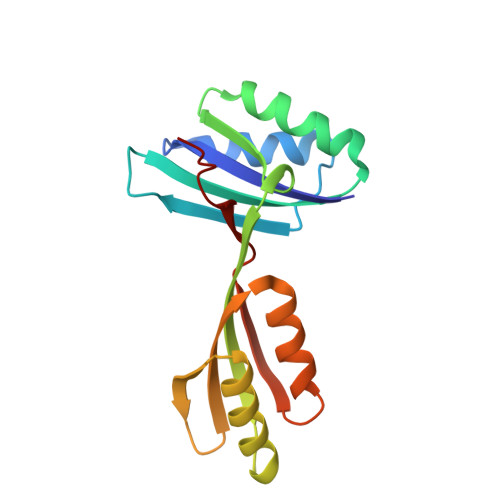
Structure of the Regulatory Subunit of Acetohydroxyacid Synthase Isozyme III from Escherichia coli.
Kaplun, A., Vyazmensky, M., Zherdev, Y., Belenky, I., Slutzker, A., Mendel, S., Barak, Z., Chipman, D.M., Shaanan, B.(2006) J Mol Biology 357: 951-963
- PubMed: 16458324 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.12.077
- Primary Citation Related Structures:
2F1F - PubMed Abstract:
The enzyme acetohydroxyacid synthase (AHAS) catalyses the first common step in the biosynthesis of the three branched-chain amino acids. Enzymes in the AHAS family generally consist of regulatory and catalytic subunits. Here, we describe the first crystal structure of an AHAS regulatory subunit, the ilvH polypeptide, determined at a resolution of 1.75 A. IlvH is the regulatory subunit of one of three AHAS isozymes expressed in Escherichia coli, AHAS III. The protein is a dimer, with two beta alpha beta beta alpha beta ferredoxin domains in each monomer. The two N-terminal domains assemble to form an ACT domain structure remarkably close to the one predicted by us on the basis of the regulatory domain of 3-phosphoglycerate dehydrogenase (3PGDH). The two C-terminal domains combine so that their beta-sheets are roughly positioned back-to-back and perpendicular to the extended beta-sheet of the N-terminal ACT domain. On the basis of the properties of mutants and a comparison with 3PGDH, the effector (valine) binding sites can be located tentatively in two symmetrically related positions in the interface between a pair of N-terminal domains. The properties of mutants of the ilvH polypeptide outside the putative effector-binding site provide further insight into the functioning of the holoenzyme. The results of this study open avenues for further studies aimed at understanding the mechanism of regulation of AHAS by small-molecule effectors.
- Department of Life Sciences, Ben-Gurion University of the Negev, Beer-Sheva 84105, Israel.
Organizational Affiliation: