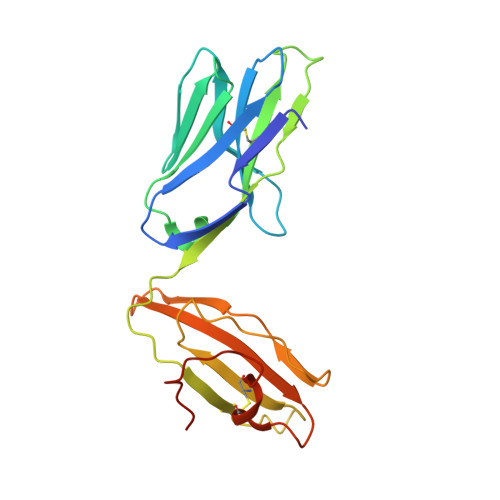
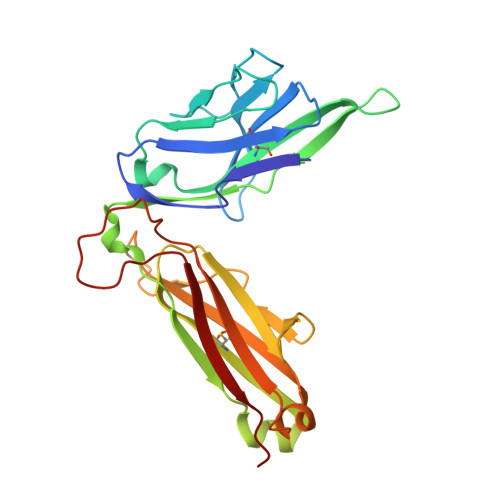
A structural basis for selection and cross-species reactivity of the semi-invariant NKT cell receptor in CD1d/glycolipid recognition
Kjer-Nielsen, L., Borg, N.A., Pellicci, D.G., Beddoe, T., Kostenko, L., Clements, C.S., Williamson, N.A., Smyth, M.J., Besra, G.S., Reid, H.H., Bharadwaj, M., Godfrey, D.I., Rossjohn, J., McCluskey, J.(2006) J Exp Medicine 203: 661-673
- PubMed: 16505140 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1084/jem.20051777
- Primary Citation Related Structures:
2EYR, 2EYS, 2EYT - PubMed Abstract:
Little is known regarding the basis for selection of the semi-invariant alphabeta T cell receptor (TCR) expressed by natural killer T (NKT) cells or how this mediates recognition of CD1d-glycolipid complexes. We have determined the structures of two human NKT TCRs that differ in their CDR3beta composition and length. Both TCRs contain a conserved, positively charged pocket at the ligand interface that is lined by residues from the invariant TCR alpha- and semi-invariant beta-chains. The cavity is centrally located and ideally suited to interact with the exposed glycosyl head group of glycolipid antigens. Sequences common to mouse and human invariant NKT TCRs reveal a contiguous conserved "hot spot" that provides a basis for the reactivity of NKT cells across species. Structural and functional data suggest that the CDR3beta loop provides a plasticity mechanism that accommodates recognition of a variety of glycolipid antigens presented by CD1d. We propose a model of NKT TCR-CD1d-glycolipid interaction in which the invariant CDR3alpha loop is predicted to play a major role in determining the inherent bias toward CD1d. The findings define a structural basis for the selection of the semi-invariant alphabeta TCR and the unique antigen specificity of NKT cells.
- Department of Microbiology and Immunology, University of Melbourne, Parkville, Victoria 3010, Australia.
Organizational Affiliation: