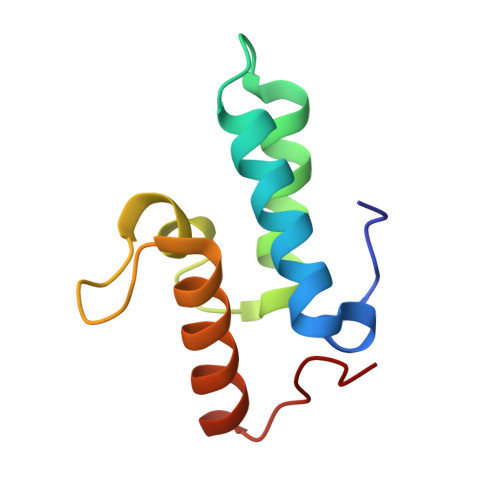
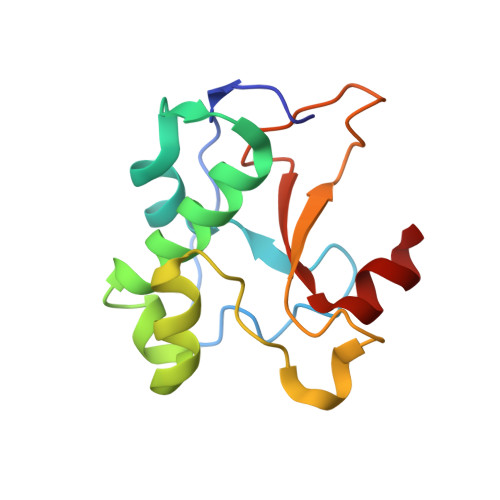
Computational Design of a New Hydrogen Bond Network and at Least a 300-fold Specificity Switch at a Protein-Protein Interface.
Joachimiak, L.A., Kortemme, T., Stoddard, B.L., Baker, D.(2006) J Mol Biology 361: 195-208
- PubMed: 16831445 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.05.022
- Primary Citation Related Structures:
2ERH - PubMed Abstract:
The redesign of protein-protein interactions is a stringent test of our understanding of molecular recognition and specificity. Previously we engineered a modest specificity switch into the colicin E7 DNase-Im7 immunity protein complex by identifying mutations that are disruptive in the native complex, but can be compensated by mutations on the interacting partner. Here we extend the approach by systematically sampling alternate rigid body orientations to optimize the interactions in a binding mode specific manner. Using this protocol we designed a de novo hydrogen bond network at the DNase-immunity protein interface and confirmed the design with X-ray crystallographic analysis. Subsequent design of the second shell of interactions guided by insights from the crystal structure on tightly bound water molecules, conformational strain, and packing defects yielded new binding partners that exhibited specificities of at least 300-fold between the cognate and the non-cognate complexes. This multi-step approach should be applicable to the design of polar protein-protein interactions and contribute to the re-engineering of regulatory networks mediated by protein-protein interactions.
- Howard Hughes Medical Institute & Department of Biochemistry, University of Washington, Seattle, 98195-7350, USA.
Organizational Affiliation: