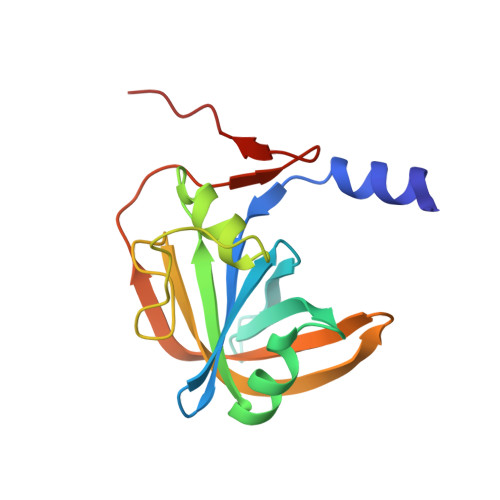
Crystal structure of the flavin reductase component (HpaC) of 4-hydroxyphenylacetate 3-monooxygenase from Thermus thermophilus HB8: Structural basis for the flavin affinity
Kim, S.H., Hisano, T., Iwasaki, W., Ebihara, A., Miki, K.(2008) Proteins 70: 718-730
- PubMed: 17729270 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21534
- Primary Citation Related Structures:
2ECR, 2ECU, 2ED4 - PubMed Abstract:
The two-component enzyme, 4-hydroxyphenylacetate 3-monooxygenase, catalyzes the conversion of 4-hydroxyphenylacetate to 3,4-dihydroxyphenylacetate. In the overall reaction, the oxygenase component (HpaB) introduces a hydroxyl group into the benzene ring of 4-hydroxyphenylacetate using molecular oxygen and reduced flavin, while the reductase component (HpaC) provides free reduced flavins for HpaB. The crystal structures of HpaC from Thermus thermophilus HB8 in the ligand-free form, the FAD-containing form, and the ternary complex with FAD and NAD(+) were determined. In the ligand-free form, two large grooves are present at the dimer interface, and are occupied by water molecules. A structural analysis of HpaC containing FAD revealed that FAD has a low occupancy, indicating that it is not tightly bound to HpaC. This was further confirmed in flavin dissociation experiments, showing that FAD can be released from HpaC. The structure of the ternary complex revealed that FAD and NAD(+) are bound in the groove in the extended and folded conformation, respectively. The nicotinamide ring of NAD(+) is sandwiched between the adenine ring of NAD(+) and the isoalloxazine ring of FAD. The distance between N5 of the isoalloxazine ring and C4 of the nicotinamide ring is about 3.3 A, sufficient to permit hydride transfer. The structures of these three states are essentially identical, however, the side chains of several residues show small conformational changes, indicating an induced fit upon binding of NADH. Inactivity with respect to NADPH can be explained as instability of the binding of NADPH with the negatively charged 2'-phosphate group buried inside the complex, as well as a possible repulsive effect by the dipole of helix alpha1. A comparison of the binding mode of FAD with that in PheA2 from Bacillus thermoglucosidasius A7, which contains FAD as a prosthetic group, reveals remarkable conformational differences in a less conserved loop region (Gly83-Gly94) involved in the binding of the AMP moiety of FAD. These data suggest that variations in the affinities for FAD in the reductases of the two-component flavin-diffusible monooxygenase family may be attributed to difference in the interaction between the AMP moiety of FAD and the less conserved loop region which possibly shows structural divergence.
- SPring-8 Center, RIKEN Harima Institute, Koto 1-1-1, Sayo-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: