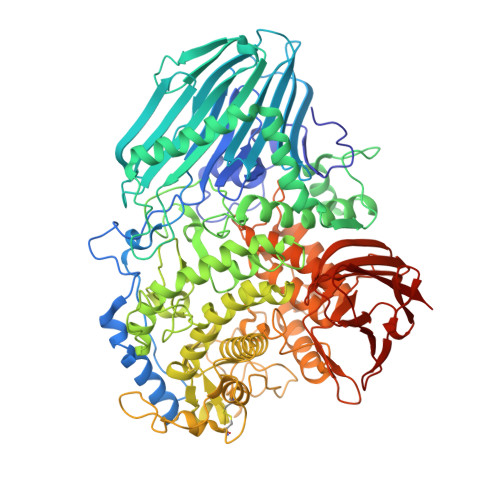
Structural basis on the catalytic reaction mechanism of novel 1,2-alpha-L-fucosidase (AFCA) from Bifidobacterium bifidum
Nagae, M., Tsuchiya, A., Katayama, T., Yamamoto, K., Wakatsuki, S., Kato, R.(2007) J Biological Chem 282: 18497-18509
- PubMed: 17459873 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M702246200
- Primary Citation Related Structures:
2EAB, 2EAC, 2EAD, 2EAE - PubMed Abstract:
1,2-alpha-L-fucosidase (AfcA), which hydrolyzes the glycosidic linkage of Fucalpha1-2Gal via an inverting mechanism, was recently isolated from Bifidobacterium bifidum and classified as the first member of the novel glycoside hydrolase family 95. To better understand the molecular mechanism of this enzyme, we determined the x-ray crystal structures of the AfcA catalytic (Fuc) domain in unliganded and complexed forms with deoxyfuconojirimycin (inhibitor), 2'-fucosyllactose (substrate), and L-fucose and lactose (products) at 1.12-2.10 A resolution. The AfcA Fuc domain is composed of four regions, an N-terminal beta region, a helical linker, an (alpha/alpha)6 helical barrel domain, and a C-terminal beta region, and this arrangement is similar to bacterial phosphorylases. In the complex structures, the ligands were buried in the central cavity of the helical barrel domain. Structural analyses in combination with mutational experiments revealed that the highly conserved Glu566 probably acts as a general acid catalyst. However, no carboxylic acid residue is found at the appropriate position for a general base catalyst. Instead, a water molecule stabilized by Asn423 in the substrate-bound complex is suitably located to perform a nucleophilic attack on the C1 atom of L-fucose moiety in 2'-fucosyllactose, and its location is nearly identical near the O1 atom of beta-L-fucose in the products-bound complex. Based on these data, we propose and discuss a novel catalytic reaction mechanism of AfcA.
- Structural Biology Research Center, Photon Factory, Insititute of Materials Structure Science, High Energy Accelerator Research Organization (KEK), Tsukuba, Ibaraki 305-0801.
Organizational Affiliation: