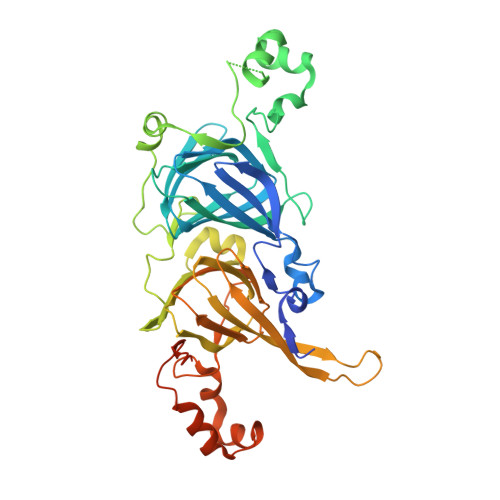
Characterization and crystallography of recombinant 7S globulins of Adzuki bean and structure-function relationships with 7S globulins of various crops.
Fukuda, T., Maruyama, N., Salleh, M.R., Mikami, B., Utsumi, S.(2008) J Agric Food Chem 56: 4145-4153
- PubMed: 18461964
- DOI: https://doi.org/10.1021/jf072667b
- Primary Citation of Related Structures:
2EA7, 2EAA - PubMed Abstract:
The recombinant proteins Adzuki 7S1, Adzuki 7S2, and Adzuki 7S3 were prepared through the Escherichia coli expression systems of three kinds of adzuki bean cDNAs. The recombinant proteins exhibited intrinsic thermal stabilities, surface hydrophobicities, and solubilities, although the homology of their amino acid sequences ranged from 95-98%. To understand why these individual proteins exhibited different properties, their three-dimensional structures were elucidated. The three proteins were successfully crystallized, and the three-dimensional structures of Adzuki 7S1 and Adzuki 7S3 were determined. The properties and structures of these two proteins were comprehensively compared with those of recombinant 7S globulins (soybean beta-conglycinins beta and alpha'c and mungbean 8Salpha) reported previously. It was likely that cavity sizes, hydrogen bonds, salt bridges, hydrophobic interactions, and lengths of loops determine the thermal stabilities of 7S globulins, and results indicated that cavity sizes strongly contribute to such stability. Surface hydrophobicity was also found to be determined not only by distributions of hydrophobic residues on the molecular surface. Furthermore, solubility at neutral and weak alkaline pH values at mu = 0.08 was found to be dominantly influenced by the electrostatic surface potentials.
- Laboratory of Food Quality Design and Development, Graduate School of Agriculture, Kyoto University, Kyoto, Japan.
Organizational Affiliation: