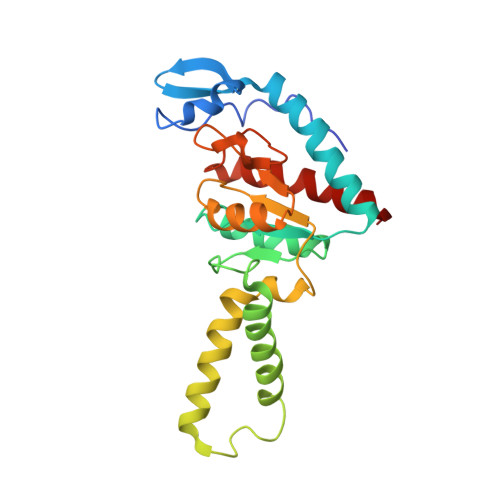
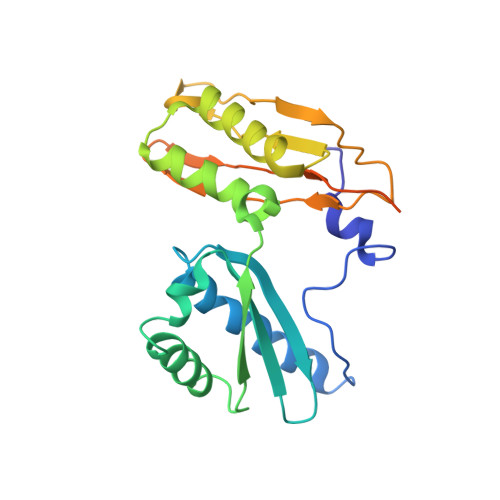
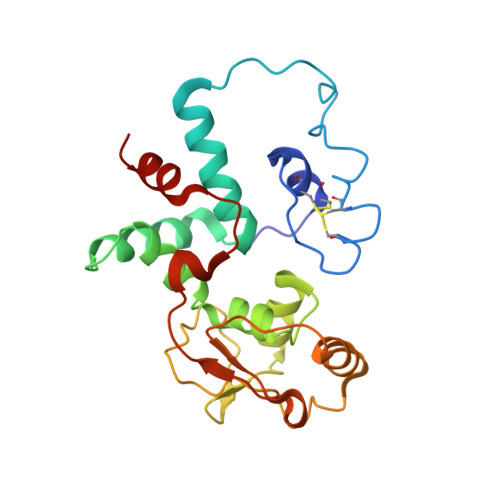
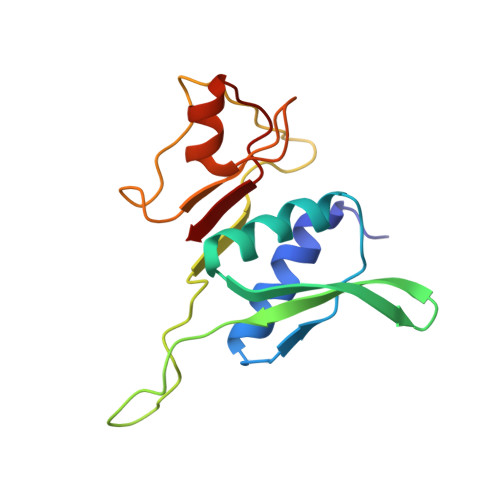
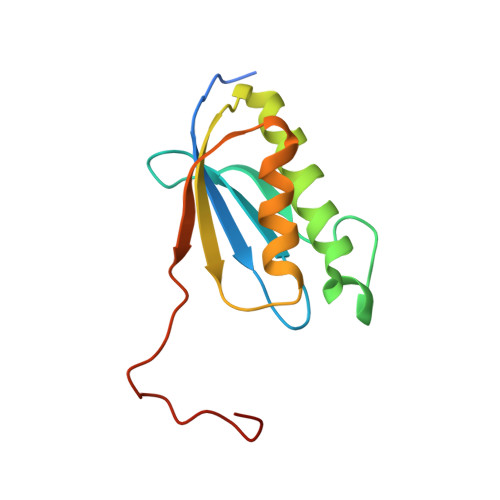
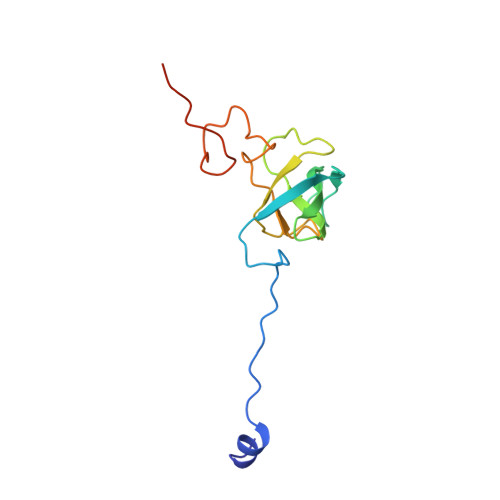
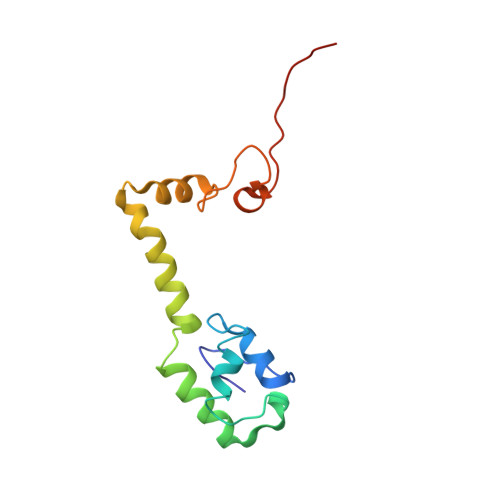
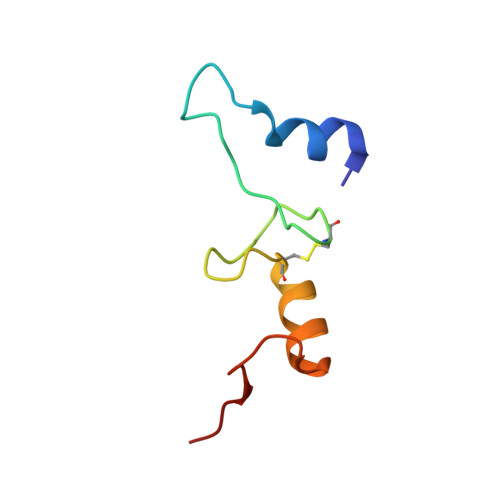
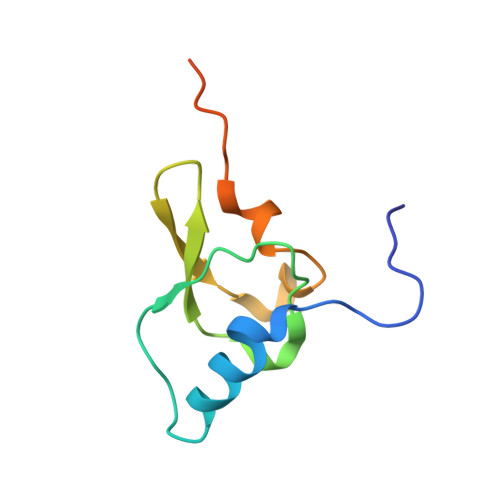
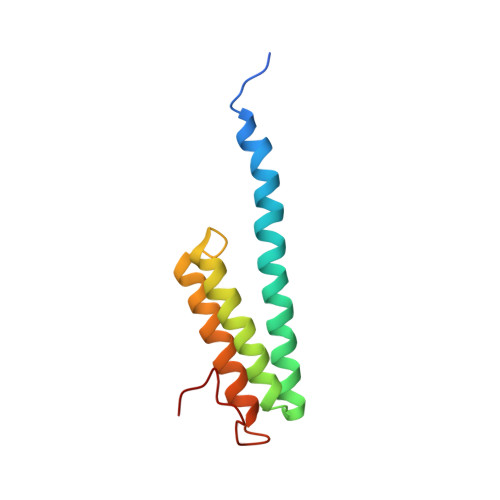
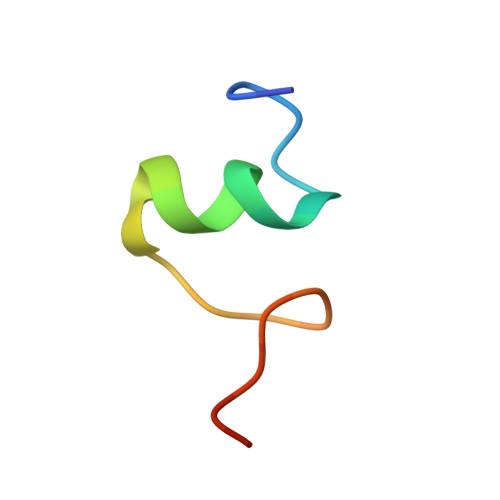
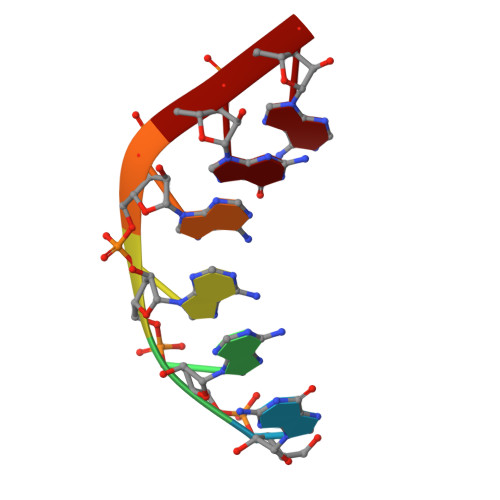
A snapshot of the 30S ribosomal subunit capturing mRNA via the Shine-Dalgarno interaction
Kaminishi, T., Wilson, D.N., Takemoto, C., Harms, J.M., Kawazoe, M., Schluenzen, F., Hanawa-Suetsugu, K., Shirouzu, M., Fucini, P., Yokoyama, S.(2007) Structure 15: 289-297
- PubMed: 17355865
- DOI: https://doi.org/10.1016/j.str.2006.12.008
- Primary Citation of Related Structures:
2E5L - PubMed Abstract:
In the initiation phase of bacterial translation, the 30S ribosomal subunit captures mRNA in preparation for binding with initiator tRNA. The purine-rich Shine-Dalgarno (SD) sequence, in the 5' untranslated region of the mRNA, anchors the 30S subunit near the start codon, via base pairing with an anti-SD (aSD) sequence at the 3' terminus of 16S rRNA. Here, we present the 3.3 A crystal structure of the Thermus thermophilus 30S subunit bound with an mRNA mimic. The duplex formed by the SD and aSD sequences is snugly docked in a "chamber" between the head and platform domains, demonstrating how the 30S subunit captures and stabilizes the otherwise labile SD helix. This location of the SD helix is suitable for the placement of the start codon AUG in the immediate vicinity of the mRNA channel, in agreement with reported crosslinks between the second position of the start codon and G1530 of 16S rRNA.
- RIKEN Genomic Sciences Center, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: