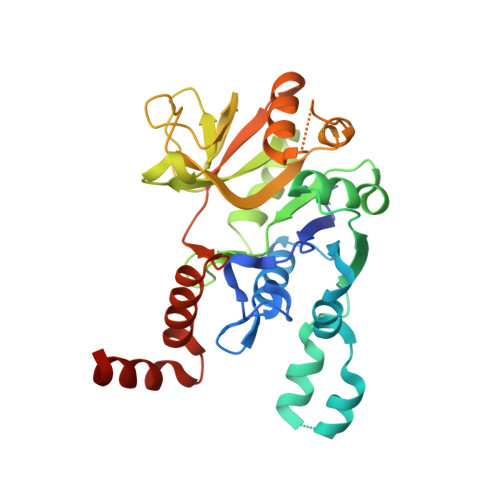
The molecular architecture of glucose-1-phosphate uridylyltransferase
Thoden, J.B., Holden, H.M.(2007) Protein Sci 16: 432-440
- PubMed: 17322528
- DOI: https://doi.org/10.1110/ps.062626007
- Primary Citation Related Structures:
2E3D - PubMed Abstract:
Glucose-1-phosphate uridylyltransferase, also referred to as UDP-glucose pyrophosphorylase or UGPase, catalyzes the formation of UDP-glucose from glucose-1-phosphate and UTP. Not surprisingly, given the central role of UDP-glucose in glycogen synthesis and in the production of glycolipids, glycoproteins, and proteoglycans, the enzyme is ubiquitous in nature. Interestingly, however, the prokaryotic and eukaryotic forms of the enzyme are unrelated in amino acid sequence and structure. Here we describe the cloning and structural analysis to 1.9 A resolution of the UGPase from Escherichia coli. The protein is a tetramer with 222 point group symmetry. Each subunit of the tetramer is dominated by an eight-stranded mixed beta-sheet. There are two additional layers of beta-sheet (two and three strands) and 10 alpha-helices. The overall fold of the molecule is remarkably similar to that observed for glucose-1-phosphate thymidylyltransferase in complex with its product, dTDP-glucose. On the basis of this similarity, a UDP-glucose moiety has been positioned into the active site of UGPase. This protein/product model predicts that the side chains of Gln 109 and Asp 137, respectively, serve to anchor the uracil ring and the ribose of UDP-glucose to the protein. The beta-phosphoryl group of the product is predicted to lie within hydrogen bonding distance to the epsilon-nitrogen of Lys 202 whereas the carboxylate group of Glu 201 is predicted to bridge the 2'- and 3'-hydroxyl groups of the glucosyl moiety. Details concerning the overall structure of UGPase and a comparison with glucose-1-phosphate thymidylyltransferase are presented.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: