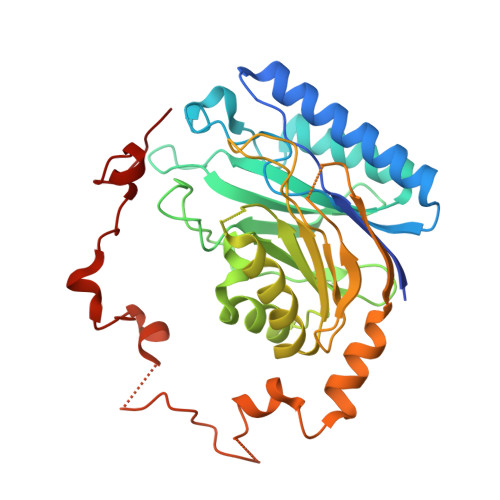
Crystal structure of Helicobacter pylori formamidase AmiF reveals a cysteine-glutamate-lysine catalytic triad
Hung, C.-L., Liu, J.-H., Chiu, W.-C., Huang, S.-W., Hwang, J.-K., Wang, W.-C.(2007) J Biological Chem 282: 12220-12229
- PubMed: 17307742 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M609134200
- Primary Citation Related Structures:
2DYU, 2DYV, 2E2K, 2E2L - PubMed Abstract:
Helicobacter pylori AmiF formamidase that hydrolyzes formamide to produce formic acid and ammonia belongs to a member of the nitrilase superfamily. The crystal structure of AmiF was solved to 1.75A resolution using single-wavelength anomalous dispersion methods. The structure consists of a homohexamer related by 3-fold symmetry in which each subunit has an alpha-beta-beta-alpha four-layer architecture characteristic of the nitrilase superfamily. One exterior alpha layer faces the solvent, whereas the other one associates with that of the neighbor subunit, forming a tight alpha-beta-beta-alpha-alpha-beta-beta-alpha dimer. The apo and liganded crystal structures of an inactive mutant C166S were also determined to 2.50 and 2.30 A, respectively. These structures reveal a small formamide-binding pocket that includes Cys(166), Glu(60), and Lys(133) catalytic residues, in which Cys(166) acts as a nucleophile. Analysis of the liganded AmiF and N-carbamoyl d-amino acid amidohydrolase binding pockets reveals a common Cys-Glu-Lys triad, another conserved glutamate, and different subsets of ligand-binding residues. Molecular dynamic simulations show that the conserved triad has minimal fluctuations, catalyzing the hydrolysis of a specific nitrile or amide in the nitrilase superfamily efficiently.
- Institute of Molecular and Cellular Biology and Department of Life Science, National Tsing Hua University, Hsinchu 300, Taiwan.
Organizational Affiliation: