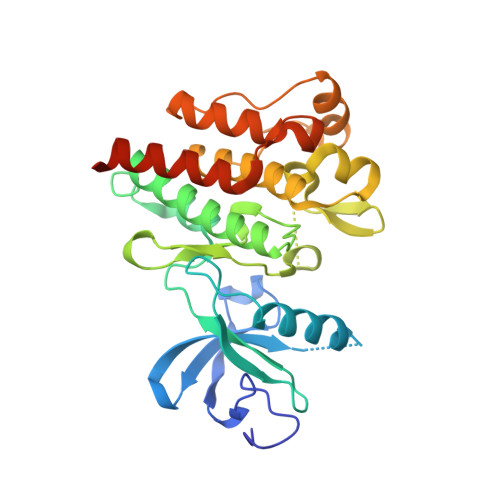
Structural factors contributing to the Abl/Lyn dual inhibitory activity of 3-substituted benzamide derivatives
Horio, T., Hamasaki, T., Inoue, T., Wakayama, T., Itou, S., Naito, H., Asaki, T., Hayase, H., Niwa, T.(2007) Bioorg Med Chem Lett 17: 2712-2717
- PubMed: 17376680 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.03.002
- Primary Citation Related Structures:
2E2B - PubMed Abstract:
To investigate why 3-substituted benzamide derivatives show dual inhibition of Abl and Lyn protein tyrosine kinases, we determined their inhibitory activities against Abl and Lyn, carried out molecular modeling, and conducted a structure-activity relationship study with the aid of a newly determined X-ray structure of the Abl/Lyn dual inhibitor INNO-406 (formerly known as NS-187) bound to human Abl. We found that this series of compounds interacted with both kinases in very similar ways, so that they can inhibit both kinases effectively.
- Research Laboratories, Nippon Shinyaku Co. Ltd., 3-14-1 Sakura, Tsukuba, Ibaraki 305-0003, Japan.
Organizational Affiliation: