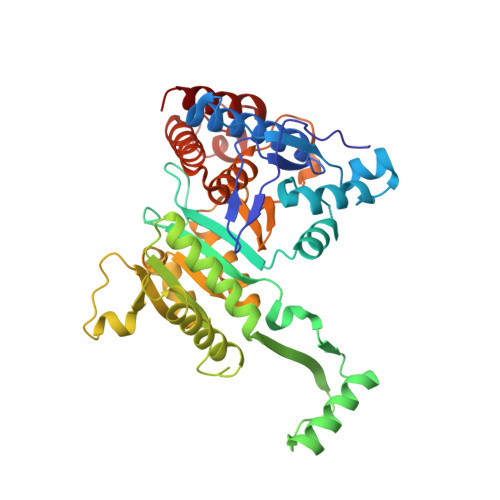
Crystal Structures of the Putative Isocitrate Dehydrogenase fromSulfolobus tokodaiiStrain 7 in the Apo and NADP+-Bound Forms.
Kondo, H., Murakami, M.(2018) Archaea 2018: 7571984-7571984
- PubMed: 30662370
- DOI: https://doi.org/10.1155/2018/7571984
- Primary Citation Related Structures:
2E0C, 2E5M - PubMed Abstract:
Isocitrate dehydrogenase is a catabolic enzyme that acts during the third step of the tricarboxylic acid cycle. The hypothetical protein ST2166 from the archaeon Sulfolobus tokodaii was isolated and crystallized. It shares high primary structure homology with prokaryotic NADP + -dependent IDHs, suggesting that these enzymes share a common enzymatic mechanism. The crystal structure of ST2166 was determined at 2.0 Å resolution in the apo form, and then the structure of the crystal soaked with NADP + was also determined at 2.4 Å resolution, which contained NADP + bound at the putative active site. Comparisons between the structures of apo and NADP + -bound forms and NADP-IDHs from other prokaryotes suggest that prokaryotic NADP-IDHs recognize their cofactors using conserved Lys335, Tyr336, and Arg386 in ST2166 at the opening cleft before the domain closure.
- Department of Physics, Graduate School of Science, Nagoya University, 464-8602 Nagoya, Japan.
Organizational Affiliation: