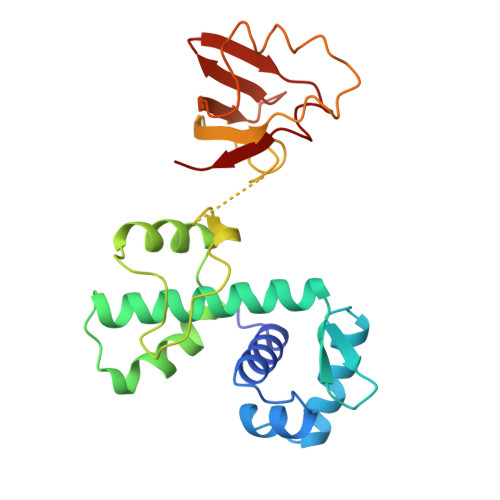
High-resolution structure of the diphtheria toxin repressor complexed with cobalt and manganese reveals an SH3-like third domain and suggests a possible role of phosphate as co-corepressor.
Qiu, X., Pohl, E., Holmes, R.K., Hol, W.G.(1996) Biochemistry 35: 12292-12302
- PubMed: 8823163 Search on PubMed
- DOI: https://doi.org/10.1021/bi960861d
- Primary Citation Related Structures:
2DTR - PubMed Abstract:
The crystal structure of diphtheria toxin repressor (DtxR) in complex with the corepressor Co2+ has been determined at 2.0 A resolution and in complex with Mn2+ at 2.2 A resolution. The structure of the flexible third domain could be determined at this high resolution. It appears to contain five antiparallel strands exhibiting a fold very similar to the SH3 domain. A superposition of 46 equivalent C alpha atoms of DtxR and alpha-spectrin SH3 resulted in an rms deviation of 3.0 A. The sequence identity is only 7%. This third domain of DtxR appears to have no interactions with the DNA binding domain nor with the metal binding domain of the repressor. Yet, flexibility in the region between the second and the third domain allows in principle significant conformational changes such as might occur upon DNA binding. The two metal binding sites in the second domain have been unraveled in considerable detail. Metal binding site 1 was well occupied in both the cobalt and manganese structures and showed a surprising sulfate ion as ligand. The sulfate was proven beyond doubt by the high peak at its position in a selenate versus sulfate difference Fourier. The presence of the intriguing sulfate ion at such a crucial position near the metal corepressor suggests the possibility that under physiological conditions phosphate may act as a "co-corepressor" for this class of metal-regulated DNA binding proteins in Corynebacteria, Mycobacteria, and related organisms. The second metal binding site is significantly different in these two DtxR structures. In the 2.0 A cobalt structure, the site is not occupied by a metal ion. In the 2.2 A manganese structure the site is well occupied, at approximately the same position as observed previously in cadmium DtxR. The ligands are Glu105, His106, the carbonyl oxygen of Cys102, and a water molecule. The reasons for differential occupancy of this site in different structures are intriguing and require further investigations.
- Department of Biological Structure, School of Medicine, University of Washington, Seattle 98195, USA.
Organizational Affiliation: