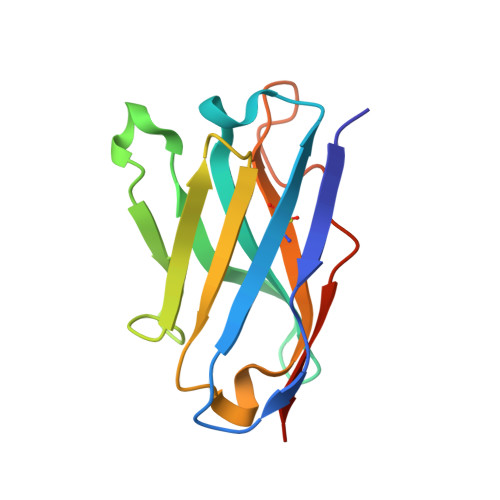
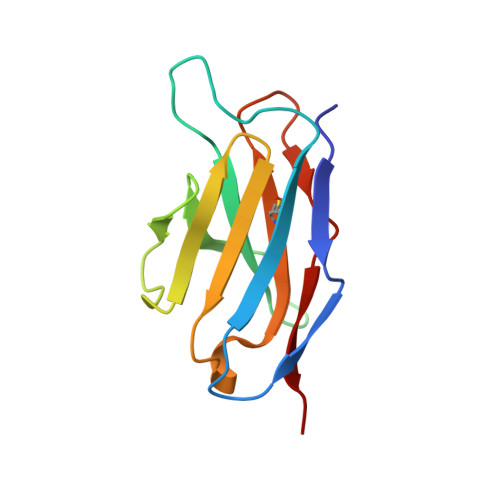
The pH-dependent structural variation of complementarity-determining region H3 in the crystal structures of the Fv fragment from an anti-dansyl monoclonal antibody.
Nakasako, M., Takahashi, H., Shimba, N., Shimada, I., Arata, Y.(1999) J Mol Biology 291: 117-134
- PubMed: 10438610 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2931
- Primary Citation Related Structures:
1DLF, 2DLF - PubMed Abstract:
The Fv fragment from an anti-dansyl antibody was optimally crystallized into two crystal forms having slightly different lattice dimensions at pH 5.25 and 6.75. The two crystal structures were determined and refined at high resolution at 112 K (at 1.45 A for the crystal at pH 5.25 and at 1.55 A for that at pH 6.75). In the two crystal structures, marked differences were identified in the first half of CDRH3 s having an amino acid sequence of Ile95H-Tyr96H-Tyr97H-His98H-Tyr99H-Pro1 00H-Trp100aH-Phe100bH-Ala101H- Tyr102H. NMR pH titration experiments revealed the p Kavalues of four histidine residues (His27dL, His93L, His55H and His98H) exposed to solvent. Only His98H (p Ka=6.3) completely changed its protonation state between the two crystallization conditions. In addition, the environmental structures including hydration water molecules around the four histidine residues were carefully compared. While the hydration structures around His27dL, His93L and His55H were almost invariant between the two crystal structures, those around His98Hs showed great difference in spite of the small conformational difference of His98H between the two crystal structures. These spectroscopic and crystallographic findings suggested that the change in the protonation state in His98H was responsible for the structural differences between pH 5.25 and 6.75. In addition, the most plausible binding site of the dansyl group was mapped into the present structural models with our previous NMR experimental results. The complementarity-determining regions H1, H3 and the N-terminal region in the VH domain formed the site. The side-chain of Tyr96H occupied the site and interacted with Phe27H of H1, giving a clue for the binding mode of the dansyl group in the site.
- Precursory Research for Embryonic Science and Technology (PRESTO), Japan Science and Technology Corporation (JST) Institute of Molecular and Cellular Biosciences, The University of Tokyo, Yayoi, Bunkyoku, Tokyo, 113-0032, Japan. nakasako@ian.u-tokyo.ac.jp
Organizational Affiliation: