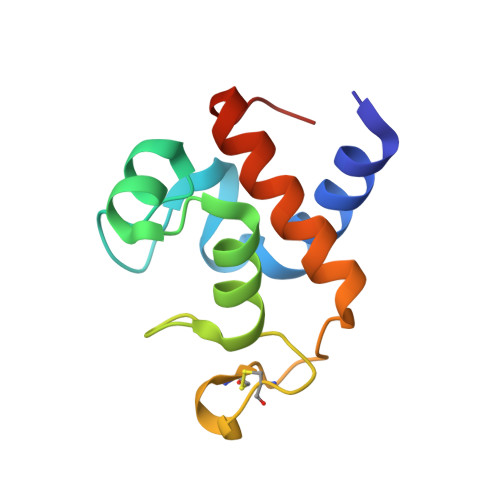
Crystal structure of oxidized cytochrome c(6A) from Arabidopsis thaliana
Chida, H., Yokoyama, T., Kawai, F., Nakazawa, A., Akazaki, H., Takayama, Y., Hirano, T., Suruga, K., Satoh, T., Yamada, S., Kawachi, R., Unzai, S., Nishio, T., Park, S.-Y., Oku, T.(2006) FEBS Lett 580: 3763-3768
- PubMed: 16777100 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2006.05.067
- Primary Citation Related Structures:
2DGE - PubMed Abstract:
Compared with algal and cyanobacterial cytochrome c(6), cytochrome c(6A) from higher plants contains an additional loop of 12 amino acid residues. We have determined the first crystal structure of cytochrome c(6A) from Arabidopsis thaliana at 1.5 Angstrom resolution in order to help elucidate its function. The overall structure of cytochrome c(6A) follows the topology of class I c-type cytochromes in which the heme prosthetic group covalently binds to Cys16 and Cys19, and the iron has octahedral coordination with His20 and Met60 as the axial ligands. Two cysteine residues (Cys67 and Cys73) within the characteristic 12 amino acids loop form a disulfide bond, contributing to the structural stability of cytochrome c(6A). Our model provides a chemical basis for the known low redox potential of cytochrome c(6A) which makes it an unsuitable electron carrier between cytochrome b(6)f and PSI.
- Bio-organic Chemistry Laboratory, Graduate School of Bioresource Sciences, Nihon University, Kameino 1866, Fujisawa-shi, Kanagawa 252-8510, Japan.
Organizational Affiliation: