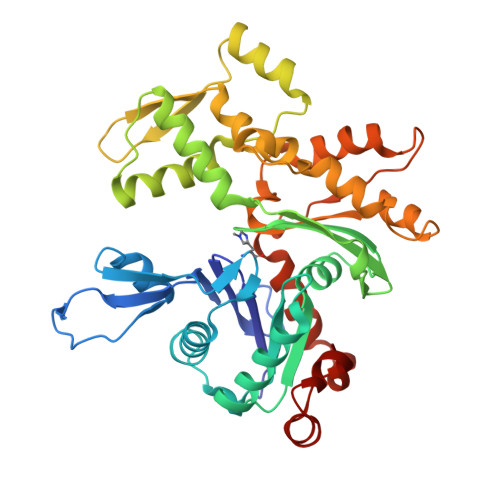
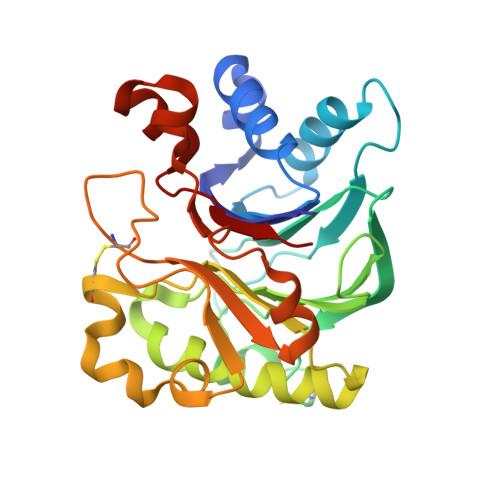
Structural basis for the actin-binding function of missing-in-metastasis
Lee, S.H., Kerff, F., Chereau, D., Ferron, F., Klug, A., Dominguez, R.(2007) Structure 15: 145-155
- PubMed: 17292833 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2006.12.005
- Primary Citation Related Structures:
2D1K, 2D1L - PubMed Abstract:
The adaptor protein missing-in-metastasis (MIM) contains independent F- and G-actin binding domains, consisting, respectively, of an N-terminal 250 aa IRSp53/MIM homology domain (IMD) and a C-terminal WASP-homology domain 2 (WH2). We determined the crystal structures of MIM's IMD and that of its WH2 bound to actin. The IMD forms a dimer, with each subunit folded as an antiparallel three-helix bundle. This fold is related to that of the BAR domain. Like the BAR domain, the IMD has been implicated in membrane binding. Yet, comparison of the structures reveals that the membrane binding surfaces of the two domains have opposite curvatures, which may determine the type of curvature of the interacting membrane. The WH2 of MIM is longer than the prototypical WH2, interacting with all four subdomains of actin. We characterize a similar WH2 at the C terminus of IRSp53 and propose that in these two proteins WH2 performs a scaffolding function.
- Department of Physiology, University of Pennsylvania School of Medicine, 3700 Hamilton Walk, Philadelphia, PA 19104, USA.
Organizational Affiliation: