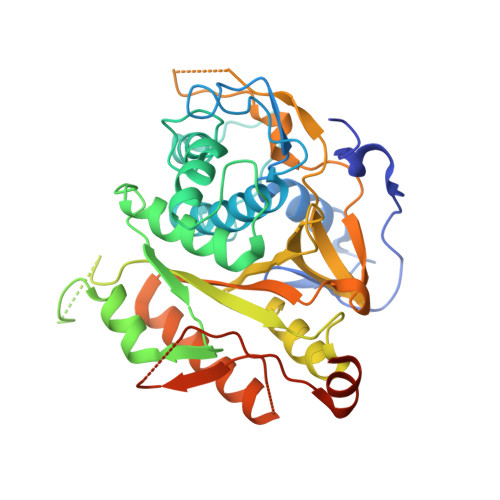
Structural Basis for the Specificity and Catalysis of Human Atg4B Responsible for Mammalian Autophagy
Sugawara, K., Suzuki, N.N., Fujioka, Y., Mizushima, N., Ohsumi, Y., Inagaki, F.(2005) J Biological Chem 280: 40058-40065
- PubMed: 16183633 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M509158200
- Primary Citation Related Structures:
2CY7 - PubMed Abstract:
Reversible modification of Atg8 with phosphatidylethanolamine is crucial for autophagy, the bulk degradation system conserved in eukaryotic cells. Atg4 is a novel cysteine protease that processes and deconjugates Atg8. Herein, we report the crystal structure of human Atg4B (HsAtg4B) at 1.9-A resolution. Despite no obvious sequence homology with known proteases, the structure of HsAtg4B shows a classical papain-like fold. In addition to the papain fold region, HsAtg4B has a small alpha/beta-fold domain. This domain is thought to be the binding site for Atg8 homologs. The active site cleft of HsAtg4B is masked by a loop (residues 259-262), implying a conformational change upon substrate binding. The structure and in vitro mutational analyses provide the basis for the specificity and catalysis of HsAtg4B. This will enable the design of Atg4-specific inhibitors that block autophagy.
- Department of Structural Biology, Graduate School of Pharmaceutical Sciences, Hokkaido University, N-12, W-6, Kita-ku, Sapporo 060-0812, Japan.
Organizational Affiliation: