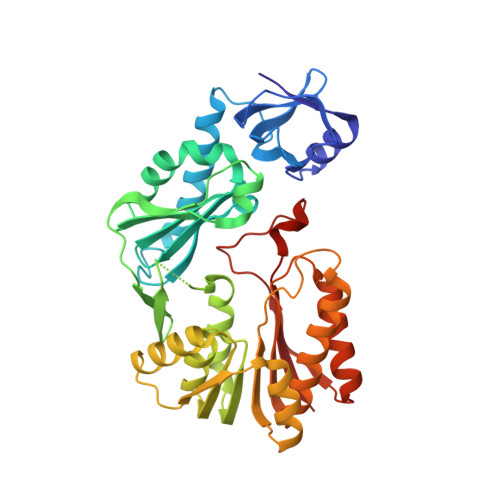
Structures of a putative RNA 5-methyluridine methyltransferase, Thermus thermophilus TTHA1280, and its complex with S-adenosyl-L-homocysteine.
Pioszak, A.A., Murayama, K., Nakagawa, N., Ebihara, A., Kuramitsu, S., Shirouzu, M., Yokoyama, S.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 867-874
- PubMed: 16511182 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309105029842
- Primary Citation Related Structures:
1WXW, 1WXX, 2CWW - PubMed Abstract:
The Thermus thermophilus hypothetical protein TTHA1280 belongs to a family of predicted S-adenosyl-L-methionine (AdoMet) dependent RNA methyltransferases (MTases) present in many bacterial and archaeal species. Inspection of amino-acid sequence motifs common to class I Rossmann-fold-like MTases suggested a specific role as an RNA 5-methyluridine MTase. Selenomethionine (SeMet) labelled and native versions of the protein were expressed, purified and crystallized. Two crystal forms of the SeMet-labelled apoprotein were obtained: SeMet-ApoI and SeMet-ApoII. Cocrystallization of the native protein with S-adenosyl-L-homocysteine (AdoHcy) yielded a third crystal form, Native-AdoHcy. The SeMet-ApoI structure was solved by the multiple anomalous dispersion method and refined at 2.55 A resolution. The SeMet-ApoII and Native-AdoHcy structures were solved by molecular replacement and refined at 1.80 and 2.60 A, respectively. TTHA1280 formed a homodimer in the crystals and in solution. Each subunit folds into a three-domain structure composed of a small N-terminal PUA domain, a central alpha/beta-domain and a C-terminal Rossmann-fold-like MTase domain. The three domains form an overall clamp-like shape, with the putative active site facing a deep cleft. The architecture of the active site is consistent with specific recognition of uridine and catalysis of methyl transfer to the 5-carbon position. The cleft is suitable in size and charge distribution for binding single-stranded RNA.
- RIKEN Genomic Sciences Center, Yokohama, Japan.
Organizational Affiliation: