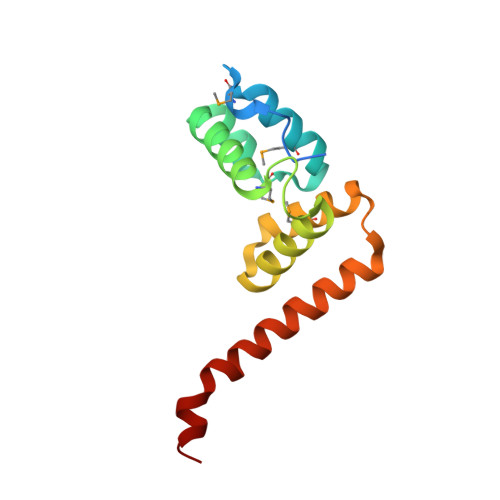
Crystal structure of the conserved protein TTHA0727 from Thermus thermophilus HB8 at 1.9 A resolution: A CMD family member distinct from carboxymuconolactone decarboxylase (CMD) and AhpD
Ito, K., Arai, R., Fusatomi, E., Kamo-Uchikubo, T., Kawaguchi, S., Akasaka, R., Terada, T., Kuramitsu, S., Shirouzu, M., Yokoyama, S.(2006) Protein Sci 15: 1187-1192
- PubMed: 16597838
- DOI: https://doi.org/10.1110/ps.062148506
- Primary Citation of Related Structures:
2CWQ - PubMed Abstract:
TTHA0727 is a conserved hypothetical protein from Thermus thermophilus HB8, with a molecular mass of 12.6 kDa. TTHA0727 belongs to the carboxymuconolactone decarboxylase (CMD) family (Pfam 02627). A sequence comparison with its homologs suggested that TTHA0727 is a distinct protein from alkylhydroperoxidase AhpD and gamma-carboxymuconolactone decarboxylase in the CMD family. Here we report the 1.9 A crystal structure of TTHA0727 (PDB ID: 2CWQ) determined by the multiwavelength anomalous dispersion method. The TTHA0727 monomer structure consists of seven alpha-helices (alpha1-alpha7) and one short 3(10)-helix. The crystal structure and the analytical ultracentrifugation revealed that TTHA0727 forms a hexameric ring structure in solution. The electrostatic potential distribution on the solvent-accessible surface of the TTHA0727 hexamer showed that positively charged regions exist on the side of the ring structure, suggesting that TTHA0727 interacts with some negatively charged molecules. A structural homology search revealed that the structure of three alpha-helices (alpha4-alpha6) is remarkably conserved, suggesting that it is the common structural motif for the CMD family proteins. In addition, the nine residues of the N-terminal tag bound to the cleft region between alpha1 and alpha3 in chains A and B of TTHA0727, implying that this region is the putative binding/active site for some small molecules.
- Protein Research Group, RIKEN Genomic Sciences Center, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: