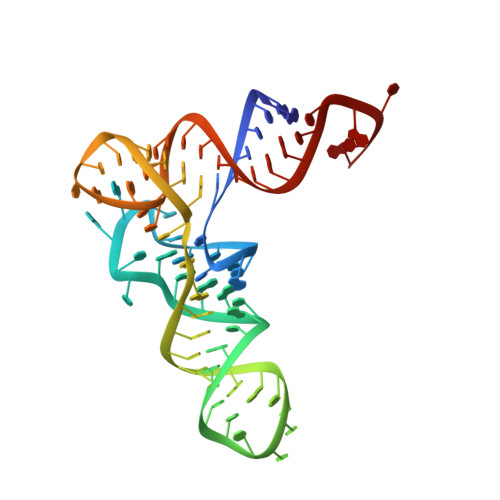
Structural bases of transfer RNA-dependent amino acid recognition and activation by glutamyl-tRNA synthetase
Sekine, S., Shichiri, M., Bernier, S., Chenevert, R., Lapointe, J., Yokoyama, S.(2006) Structure 14: 1791-1799
- PubMed: 17161369 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2006.10.005
- Primary Citation Related Structures:
2CUZ, 2CV0, 2CV2, 2DXI - PubMed Abstract:
Glutamyl-tRNA synthetase (GluRS) is one of the aminoacyl-tRNA synthetases that require the cognate tRNA for specific amino acid recognition and activation. We analyzed the role of tRNA in amino acid recognition by crystallography. In the GluRS*tRNA(Glu)*Glu structure, GluRS and tRNA(Glu) collaborate to form a highly complementary L-glutamate-binding site. This collaborative site is functional, as it is formed in the same manner in pretransition-state mimic, GluRS*tRNA(Glu)*ATP*Eol (a glutamate analog), and posttransition-state mimic, GluRS*tRNA(Glu)*ESA (a glutamyl-adenylate analog) structures. In contrast, in the GluRS*Glu structure, only GluRS forms the amino acid-binding site, which is defective and accounts for the binding of incorrect amino acids, such as D-glutamate and L-glutamine. Therefore, tRNA(Glu) is essential for formation of the completely functional binding site for L-glutamate. These structures, together with our previously described structures, reveal that tRNA plays a crucial role in accurate positioning of both L-glutamate and ATP, thus driving the amino acid activation.
- Department of Biophysics and Biochemistry, Graduate School of Science, University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: