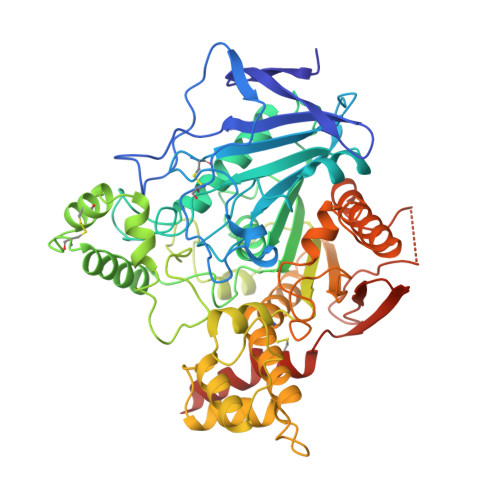
Complexes of alkylene-linked tacrine dimers with Torpedo californica acetylcholinesterase: Binding of Bis5-tacrine produces a dramatic rearrangement in the active-site gorge.
Rydberg, E.H., Brumshtein, B., Greenblatt, H.M., Wong, D.M., Shaya, D., Williams, L.D., Carlier, P.R., Pang, Y.P., Silman, I., Sussman, J.L.(2006) J Med Chem 49: 5491-5500
- PubMed: 16942022 Search on PubMed
- DOI: https://doi.org/10.1021/jm060164b
- Primary Citation Related Structures:
1ODC, 1UT6, 2CKM, 2CMF - PubMed Abstract:
The X-ray crystal structures were solved for complexes with Torpedo californica acetylcholinesterase of two bivalent tacrine derivative compounds in which the two tacrine rings were separated by 5- and 7-carbon spacers. The derivative with the 7-carbon spacer spans the length of the active-site gorge, making sandwich interactions with aromatic residues both in the catalytic anionic site (Trp84 and Phe330) at the bottom of the gorge and at the peripheral anionic site near its mouth (Tyr70 and Trp279). The derivative with the 5-carbon spacer interacts in a similar manner at the bottom of the gorge, but the shorter tether precludes a sandwich interaction at the peripheral anionic site. Although the upper tacrine group does interact with Trp279, it displaces the phenyl residue of Phe331, thus causing a major rearrangement in the Trp279-Ser291 loop. The ability of this inhibitor to induce large-scale structural changes in the active-site gorge of acetylcholinesterase has significant implications for structure-based drug design because such conformational changes in the target enzyme are difficult to predict and to model.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: