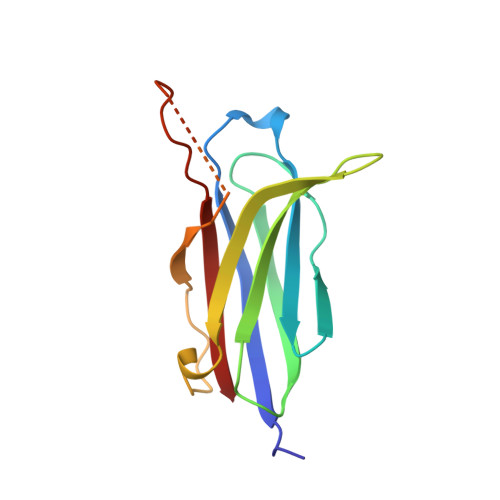
Structural Basis for a Munc13-1 Dimeric to Munc13-1/Rim Heterodimer Switch
Lu, J., Machius, M., Dulubova, I., Dai, H., Sudhof, T.C., Tomchick, D.R., Rizo, J.(2006) PLoS Biol 4: E192
- PubMed: 16732694 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.0040192
- Primary Citation Related Structures:
2CJS, 2CJT - PubMed Abstract:
C(2) domains are well characterized as Ca(2+)/phospholipid-binding modules, but little is known about how they mediate protein-protein interactions. In neurons, a Munc13-1 C(2)A-domain/RIM zinc-finger domain (ZF) heterodimer couples synaptic vesicle priming to presynaptic plasticity. We now show that the Munc13-1 C(2)A domain homodimerizes, and that homodimerization competes with Munc13-1/RIM heterodimerization. X-ray diffraction studies guided by nuclear magnetic resonance (NMR) experiments reveal the crystal structures of the Munc13-1 C(2)A-domain homodimer and the Munc13-1 C(2)A-domain/RIM ZF heterodimer at 1.44 A and 1.78 A resolution, respectively. The C(2)A domain adopts a beta-sandwich structure with a four-stranded concave side that mediates homodimerization, leading to the formation of an eight-stranded beta-barrel. In contrast, heterodimerization involves the bottom tip of the C(2)A-domain beta-sandwich and a C-terminal alpha-helical extension, which wrap around the RIM ZF domain. Our results describe the structural basis for a Munc13-1 homodimer-Munc13-1/RIM heterodimer switch that may be crucial for vesicle priming and presynaptic plasticity, uncovering at the same time an unexpected versatility of C(2) domains as protein-protein interaction modules, and illustrating the power of combining NMR spectroscopy and X-ray crystallography to study protein complexes.
- Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
Organizational Affiliation: