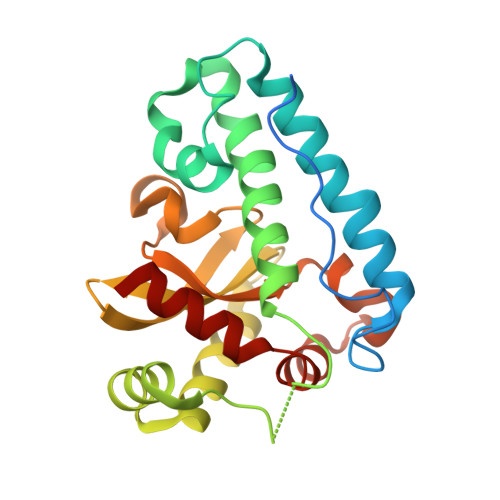
Structure of the manganese superoxide dismutase from Deinococcus radiodurans in two crystal forms.
Dennis, R.J., Micossi, E., McCarthy, J., Moe, E., Gordon, E.J., Kozielski-Stuhrmann, S., Leonard, G.A., McSweeney, S.(2006) Acta Crystallogr Sect F Struct Biol Cryst Commun 62: 325-329
- PubMed: 16582477 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309106008402
- Primary Citation Related Structures:
2CDY, 2CE4 - PubMed Abstract:
The structure of the manganese superoxide dismutase (Mn-SOD; DR1279) from Deinococcus radiodurans has been determined in two different crystal forms. Both crystal forms are monoclinic with space group P2(1). Form I has unit-cell parameters a = 44.28, b = 83.21, c = 59.52 angstroms, beta = 110.18 degrees and contains a homodimer in the asymmetric unit, with structure refinement (R = 16.8%, R(free) = 23.6%) carried out using data to d(min) = 2.2 angstroms. Form II has unit-cell parameters a = 43.57, b = 87.10, c = 116.42 angstroms, beta = 92.1 degrees and an asymmetric unit containing two Mn-SOD homodimers; structure refinement was effected to a resolution of 2.0 angstroms (R = 17.2%, R(free) = 22.3%). The resulting structures are compared with that of Mn-SOD from Escherichia coli, with which they are shown to be essentially isostructural.
- Macromolecular Crystallography Group, European Synchrotron Radiation Facility, 38043 Grenoble CEDEX 9, France.
Organizational Affiliation: