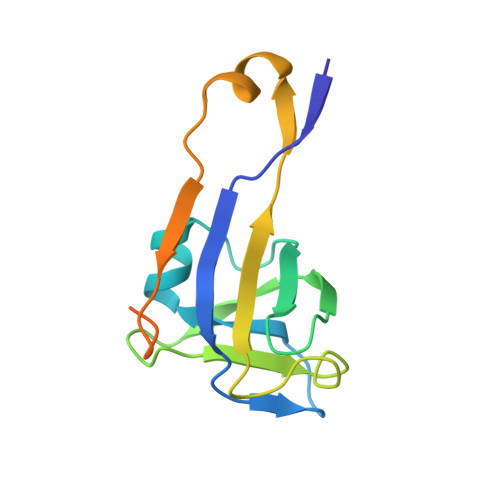
Crystal structure of uncleaved L-aspartate-alpha-decarboxylase from Mycobacterium tuberculosis.
Gopalan, G., Chopra, S., Ranganathan, A., Swaminathan, K.(2006) Proteins 65: 796-802
- PubMed: 17001646 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21126
- Primary Citation Related Structures:
2C45 - PubMed Abstract:
L-aspartate-alpha-decarboxylase (ADC) is a critical regulatory enzyme in the pantothenate biosynthetic pathway and belongs to a small class of self-cleaving and pyruvoyl-dependent amino acid decarboxylases. The expression level of ADC in Mycobacterium tuberculosis (Mtb) was confirmed by cDNA analysis, immunoblotting with an anti-ADC polyclonal antibody using whole cell lysate and immunoelectron microscopy. The recombinant ADC proenzyme from Mycobacterium tuberculosis (MtbADC) was overexpressed in E. coli and the protein structure was determined at 2.99 A resolution. The proteins fold into the double-psi beta-barrel structure. The subunits of the two tetramers (there are eight ADC molecules in the asymmetric unit) form pseudo fourfold rotational symmetry, similar to the E. coli ADC proenzyme structure. As pantothenate is synthesized in microorganisms, plants, and fungi but not in animals, structure elucidation of Mtb ADC is of substantial interest for structure-based drug development.
- Department of Biological Sciences, National University of Singapore, Singapore 117543.
Organizational Affiliation: