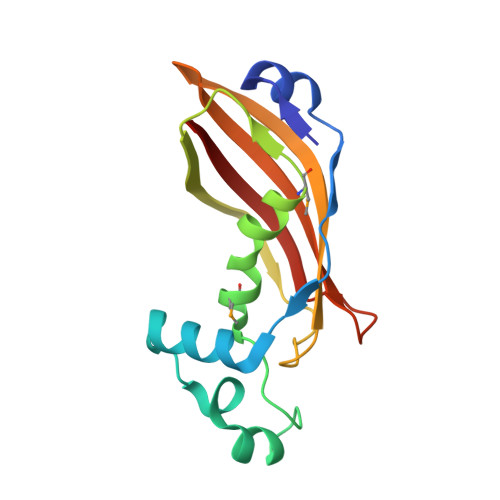
Structure and Function of Rv0130, a Conserved Hypothetical Protein from Mycobacterium Tuberculosis.
Johansson, P., Castell, A., Jones, T.A., Backbro, K.(2006) Protein Sci 15: 2300
- PubMed: 16963641
- DOI: https://doi.org/10.1110/ps.062309306
- Primary Citation Related Structures:
2C2I - PubMed Abstract:
A large fraction of the Mycobacterium tuberculosis genome codes for proteins of unknown function. We here report the structure of one of these proteins, Rv0130, solved to a resolution of 1.8 å. The Rv0130 monomer features a single hotdog fold composed of a highly curved beta-sheet on top of a long and a short alpha-helix. Two monomers in turn pack to form a double-hotdog-folded homodimer, similar to a large group of enzymes that use thiol esters as substrates. Rv0130 was found to contain a highly conserved R-specific hydratase motif buried deeply between the two monomers. Our biochemical studies show that the protein is able to hydrate a short trans-2-enoyl-coenzyme A moiety with a k(cat) of 1.1 x 10(2) sec(-1). The importance of the side chains of D40 and H45 for hydratase activity is demonstrated by site-directed mutagenesis. In contrast to many hotdog-folded proteins, a proline residue distorts the central helix of Rv0130. This distortion allows the creation of a long, curved tunnel, similar to the substrate-binding channels of long-chain eukaryotic hydratase 2 enzymes.
- Department of Cell and Molecular Biology, Uppsala University, Biomedical Center, SE-751 24 Uppsala, Sweden.
Organizational Affiliation: