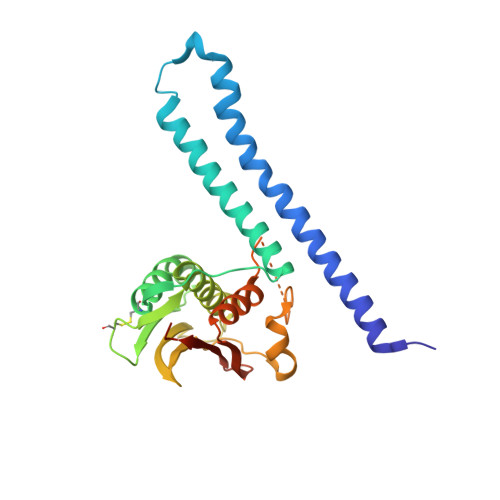
Structure of the Entire Cytoplasmic Portion of a Sensor Histidine-Kinase Protein.
Marina, A., Waldburger, C.D., Hendrickson, W.A.(2005) EMBO J 24: 4247
- PubMed: 16319927
- DOI: https://doi.org/10.1038/sj.emboj.7600886
- Primary Citation of Related Structures:
2C2A - PubMed Abstract:
The large majority of histidine kinases (HKs) are multifunctional enzymes having autokinase, phosphotransfer and phosphatase activities, and most of these are transmembrane sensor proteins. Sensor HKs possess conserved cytoplasmic phosphorylation and ATP-binding kinase domains. The different enzymatic activities require participation by one or both of these domains, implying the need for different conformational states. The catalytic domains are linked to the membrane through a coiled-coil segment that sometimes includes other domains. We describe here the first crystal structure of the complete cytoplasmic region of a sensor HK, one from the thermophile Thermotoga maritima in complex with ADPbetaN at 1.9 A resolution. The structure reveals previously unidentified functions for several conserved residues and reveals the relative disposition of domains in a state seemingly poised for phosphotransfer. The structure thereby inspires hypotheses for the mechanisms of autophosphorylation, phosphotransfer and response-regulator dephosphorylation, and for signal transduction through the coiled-coil segment. Mutational tests support the functional relevance of interdomain contacts.
- Howard Hughes Medical Institute, Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: