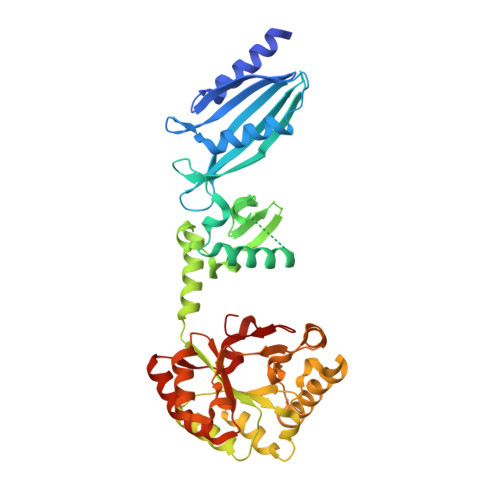
Structure and metal-dependent mechanism of peptidoglycan deacetylase, a streptococcal virulence factor.
Blair, D.E., Schuttelkopf, A.W., MacRae, J.I., van Aalten, D.M.(2005) Proc Natl Acad Sci U S A 102: 15429-15434
- PubMed: 16221761 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0504339102
- Primary Citation Related Structures:
2C1G, 2C1I - PubMed Abstract:
Streptococcus pneumoniae peptidoglycan GlcNAc deacetylase (SpPgdA) protects the Gram-positive bacterial cell wall from host lysozymes by deacetylating peptidoglycan GlcNAc residues. Deletion of the pgda gene has been shown to result in hypersensitivity to lysozyme and reduction of infectivity in a mouse model. SpPgdA is a member of the family 4 carbohydrate esterases, for which little structural information exists, and no catalytic mechanism has yet been defined. Here we describe the native crystal structure and product complexes of SpPgdA biochemical characterization and mutagenesis. The structural data show that SpPgdA is an elongated three-domain protein in the crystal. The structure, in combination with mutagenesis, shows that SpPgdA is a metalloenzyme using a His-His-Asp zinc-binding triad with a nearby aspartic acid and histidine acting as the catalytic base and acid, respectively, somewhat similar to other zinc deacetylases such as LpxC. The enzyme is able to accept GlcNAc(3) as a substrate (K(m) = 3.8 mM, k(cat) = 0.55 s(-1)), with the N-acetyl of the middle sugar being removed by the enzyme. The data described here show that SpPgdA and the other family 4 carbohydrate esterases are metalloenzymes and present a step toward identification of mechanism-based inhibitors for this important class of enzymes.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, DD1 5EH Dundee, Scotland.
Organizational Affiliation: