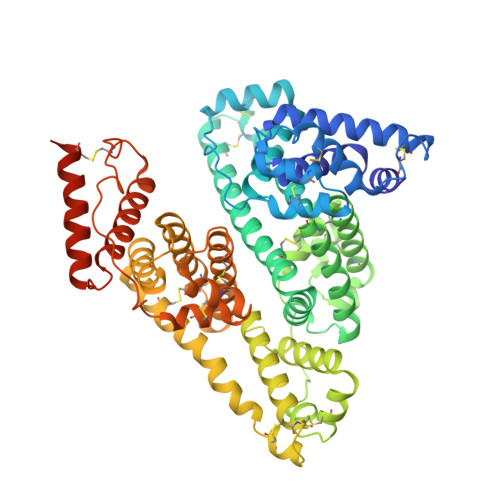
Structural Basis of the Drug-Binding Specificity of Human Serum Albumin.
Ghuman, J., Zunszain, P.A., Petitpas, I., Bhattacharya, A.A., Otagiri, M., Curry, S.(2005) J Mol Biology 353: 38
- PubMed: 16169013 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.07.075
- Primary Citation Related Structures:
2BX8, 2BXA, 2BXB, 2BXC, 2BXD, 2BXE, 2BXF, 2BXG, 2BXH, 2BXI, 2BXK, 2BXL, 2BXM, 2BXN, 2BXO, 2BXP, 2BXQ - PubMed Abstract:
Human serum albumin (HSA) is an abundant plasma protein that binds a remarkably wide range of drugs, thereby restricting their free, active concentrations. The problem of overcoming the binding affinity of lead compounds for HSA represents a major challenge in drug development. Crystallographic analysis of 17 different complexes of HSA with a wide variety of drugs and small-molecule toxins reveals the precise architecture of the two primary drug-binding sites on the protein, identifying residues that are key determinants of binding specificity and illuminating the capacity of both pockets for flexible accommodation. Numerous secondary binding sites for drugs distributed across the protein have also been identified. The binding of fatty acids, the primary physiological ligand for the protein, is shown to alter the polarity and increase the volume of drug site 1. These results clarify the interpretation of accumulated drug binding data and provide a valuable template for design efforts to modulate the interaction with HSA.
- Biophysics Section, Division of Cell and Molecular Biology, Imperial College London, South Kensington Campus, London SW7 2AZ, UK.
Organizational Affiliation: