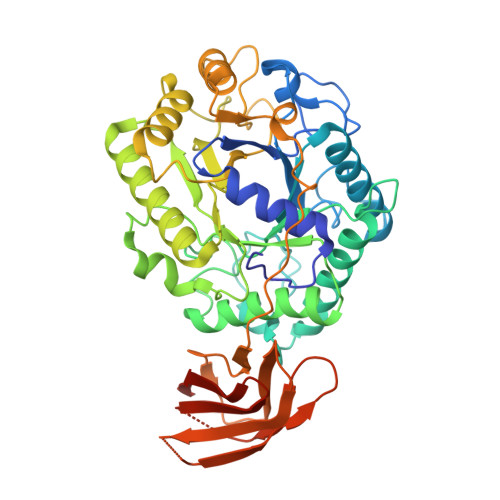
The Structure and Characterization of a Modular Endo-Beta-1,4-Mannanase from Cellulomonas Fimi
Le Nours, J., Anderson, L., Stoll, D., Stalbrand, H., Lo Leggio, L.(2005) Biochemistry 44: 12700
- PubMed: 16171384 Search on PubMed
- DOI: https://doi.org/10.1021/bi050779v
- Primary Citation Related Structures:
2BVT, 2BVY - PubMed Abstract:
The endo-beta-1,4-mannanase from the soil bacterium Cellulomonas fimi is a modular plant cell wall degrading enzyme involved in the hydrolysis of the backbone of mannan, one of the most abundant polysaccharides of the hemicellulosic network in the plant cell wall. The crystal structure of a recombinant truncated endo-beta-1,4-mannanase from C. fimi (CfMan26A-50K) was determined by X-ray crystallography to 2.25 A resolution using the molecular replacement technique. The overall structure of the enzyme consists of a core (beta/alpha)8-barrel catalytic module characteristic of clan GH-A, connected via a linker to an immunoglobulin-like module of unknown function. A complex with the oligosaccharide mannotriose to 2.9 A resolution has also been obtained. Both the native structure and the complex show a cacodylate ion bound at the -1 subsite, while subsites -2, -3, and -4 are occupied by mannotriose in the complex. Enzyme kinetic analysis and the analysis of hydrolysis products from manno-oligosaccharides and mannopentitol suggest five important active-site cleft subsites. CfMan26A-50K has a high affinity -3 subsite with Phe325 as an aromatic platform, which explains the mannose releasing property of the enzyme. Structural differences with the homologous Cellvibrio japonicus beta-1,4-mannanase (CjMan26A) at the -2 and -3 subsites may explain the poor performance of CfMan26A mutants as "glycosynthases".
- Centre for Crystallographic Studies, Biophysical Chemistry Group, Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen, Denmark.
Organizational Affiliation: