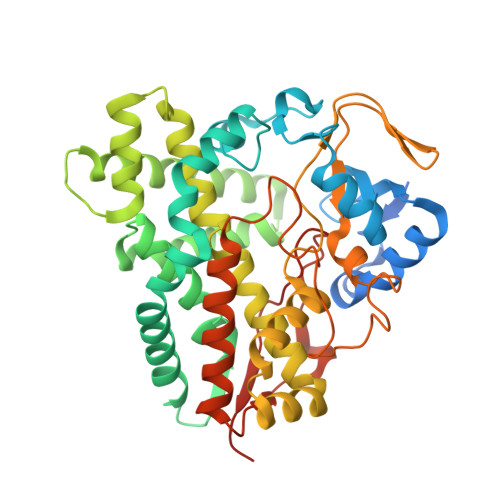
The Structural Basis for Substrate Anchoring, Active Site Selectivity, and Product Formation by P450 Pikc from Streptomyces Venezuelae.
Sherman, D.H., Li, S., Yermalitskaya, L.V., Kim, Y., Smith, J.A., Waterman, M.R., Podust, L.M.(2006) J Biological Chem 281: 26289
- PubMed: 16825192 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M605478200
- Primary Citation Related Structures:
2BVJ, 2C6H, 2C7X, 2CD8 - PubMed Abstract:
The pikromycin (Pik)/methymycin biosynthetic pathway of Streptomyces venezuelae represents a valuable system for dissecting the fundamental mechanisms of modular polyketide biosynthesis, aminodeoxysugar assembly, glycosyltransfer, and hydroxylation leading to the production of a series of macrolide antibiotics, including the natural ketolides narbomycin and pikromycin. In this study, we describe four x-ray crystal structures and allied functional studies for PikC, the remarkable P450 monooxygenase responsible for production of a number of related macrolide products from the Pik pathway. The results provide important new insights into the structural basis for the C10/C12 and C12/C14 hydroxylation patterns for the 12-(YC-17) and 14-membered ring (narbomycin) macrolides, respectively. This includes two different ligand-free structures in an asymmetric unit (resolution 2.1 A) and two co-crystal structures with bound endogenous substrates YC-17 (resolution 2.35 A)or narbomycin (resolution 1.7 A). A central feature of the enzyme-substrate interaction involves anchoring of the desosamine residue in two alternative binding pockets based on a series of distinct amino acid residues that form a salt bridge and a hydrogen-bonding network with the deoxysugar C3' dimethylamino group. Functional significance of the salt bridge was corroborated by site-directed mutagenesis that revealed a key role for Glu-94 in YC-17 binding and Glu-85 for narbomycin binding. Taken together, the x-ray structure analysis, site-directed mutagenesis, and corresponding product distribution studies reveal that PikC substrate tolerance and product diversity result from a combination of alternative anchoring modes rather than an induced fit mechanism.
- Life Sciences Institute and Department of Medicinal Chemistry, University of Michigan, Ann Arbor, Michigan, 48109, USA. davidhs@umich.edu
Organizational Affiliation: